Congressional Justification FY 2007
Department of Health and Human Services
National Institutes of Health
National Library of Medicine
2007 Congressional Justification (PDF)
FY 2007 Budget
- Organization chart
- Appropriation language
- Amounts available for obligation
- Justification narrative
- Budget mechanism table
- Budget authority by activity
- Summary of changes
- Budget authority by object class
- Salaries and expenses
- Significant items in the House, Senate and Conference Appropriations Committee Reports
- Authorizing legislation
- Appropriation history
- Detail of full-time equivalent employment (FTE)
- Detail of positions
- New positions requested
Organizational Structure
- OFFICE OF THE DIRECTOR
Donald A.B. Lindberg, M.D., Director
Betsy L. Humphreys, Deputy Director
Donald W. King, M.D., Deputy Director for Research and Education
Todd D. Danielson, Executive Officer- Division of Extramural Programs
Milton Corn, M.D., Associate Director - Division of Library Operations
Becky Lyon, Acting Associate Director - Lister Hill National Center for Biomedical Communications
Donald W. King, M.D., Acting Director - Division of Specialized Information Services
Jack W. Snyder, M.D., Ph.D., Associate Director - National Center for Biotechnology Information
David J. Lipman, M.D., Director
- Division of Extramural Programs
Appropriation Language
For carrying out section 301 and title IV of the Public Health Service Act with respect to health information communications, [$314,910,000] $313,269,000, of which $4,000,000 shall be available until expended for improvement of information systems: Provided, That in fiscal year [2006] 2007 the Library may enter into personal services contracts for the provision of services in facilities owned, operated or constructed under the jurisdiction of the National Institutes of Health. Provided further, That in addition to amounts provided herein, $8,200,000 shall be available from amounts available under section 241 of the Public Health Service Act to carry out National Information Center on Health Services Research and Health Care Technology and related health services.
[Department of Health and Human Services Appropriations Act, 2006]
Amounts Available for Obligation 1
| Source of Funding | FY 2005 Actual | FY 2006 Appropriation |
FY 2007 Estimate |
|---|---|---|---|
| Appropriation | $317,947,000 | $318,091,000 | $313,269,000 |
| Enacted Rescission | (2,801,000) | (3,181,000) | 0 |
| Subtotal, adjusted appropriation | 315,146,000 | 314,910,000 | 313,269,000 |
| Real Transfer under NIH Director's one-percent transfer authority for Roadmap | (1,992,000) | (2,814,000) | 0 |
| Comparative transfer from OD for NIH Roadmap | 1,992,000 | 2,814,000 | 0 |
| Subtotal, adjusted budget authority | 315,146,000 | 314,910,000 | 313,269,000 |
| Unobligated balance, start of year | 0 | 623 | 0 |
| Revenue from Breast Cancer Stamp 2/ | 0 | 0 | 0 |
| Unobligated balance, end of year | (623) | 0 | 0 |
| Subtotal, adjusted budget authority | 315,145,377 | 314,910,623 | 313,269,000 |
| Unobligated balance lapsing | (174,000) | 0 | 0 |
| Total obligations | 314,971,377 | 314,910,623 | 313,269,000 |
1/ Excludes the following amounts for reimbursable activities carried out by this account:
FY 2005 - $18,451,000; FY 2006 - $18,478,000; FY 2007 - $18,478,000
Justification
Authorizing Legislation: Section 301 and title IV of the Public Health Service Act, as amended.
| Budget Authority | FY 2005 Actual |
FY 2006 Appropriation |
FY 2007 Estimate |
Increase or Decrease |
|---|---|---|---|---|
| FTE | 661 | 653 | 656 | 3 |
| BA | $315,146,000 | $314,910,000 | $313,269,000 | -$1,641,000 |
This document provides justification for the Fiscal Year 2007 activities of the National Library of Medicine, including HIV/AIDS activities. A more detailed description of NIH-wide Fiscal Year 2007 HIV/AIDS activities can be found in the NIH section entitled "Office of AIDS Research (OAR)." Detailed information on the NIH Roadmap for Medical Research may be found in the Overview section.
INTRODUCTION
The National Library of Medicine (NLM) is an institution that has the potential to touch the everyday lives of all Americans. Its databases and information resources are consulted almost one billion times a year by the public and scientists alike, making it the largest purveyor of science-based health information in the world.
- Researchers and health professionals have instantaneous access to more than 15 million scientific articles on virtually any medical subject via the PubMed/Medline® database.
- The general public consults MedlinePlus®, which has consumer friendly information from the National Institutes of Health and other trusted sources.
- Those who have more specialized needs have access to the Library's databases about clinical trials, the environment, the history of the health sciences, and genomic information and molecular sequence data.
- There is also a series of information resources for special populations—American Indians, Asian Americans, those who live in the Arctic, the elderly, and those whose primary language is Spanish.
The National Library of Medicine makes all this information—and much more—accessible through the World Wide Web free and without registration.
Although we live in an increasingly "virtual" world, the Library takes great pride in collecting and preserving the literature of the biosciences. It has more than 7 million books, journals, and other forms of the literature on its shelves, making it the largest medical library in the world. While many millions use our Web resources, many thousands actually visit the Library in Bethesda, Maryland. There they have access not just to the electronic databases and contemporary medical literature, but to one of the world's greatest treasure troves of books, manuscripts, and art relating to the history of the health sciences. Used not only by scholars, these materials are frequently integrated into fascinating exhibitions and displays for visitors. Traveling versions of NLM exhibitions attract crowds across the country.
NLM's key partner in making information available is the National Network of Libraries of Medicine®. The network consists of 5,500 member institutions, including eight Regional Medical Libraries that receive NLM support, 125 resource libraries connected to medical schools, and more than 5,000 libraries located primarily in hospitals and clinics. This network is of inestimable value in assisting NLM to reach out to the wider community. They hold workshops and provide backup services for public libraries and other community organizations, demonstrate NLM databases to the public, and exhibit at meetings and conventions on behalf of NLM, thus providing the personal element that can be so important in connecting Americans to high quality information.
Several illustrations show just how applicable the Library's services can be to contemporary life. Within days after the 9/11 terrorist attack and Hurricane Katrina disaster, NLM experts had reviewed the available resources and created Web sites that provided information for both health professionals and the public in a useful form. They have done the same with avian or bird flu. The Library has several programs of grant assistance to focus the attention of informatics researchers on disaster management issues, and the Library has supported highly praised work on advanced biosurveillance techniques. Projects are also funded dealing with mental health issues, triage among multiple hospitals, and other topics relevant to natural disasters.
It is especially gratifying when NLM learns how its systems are used in real medical situations. For example, Barry Marshall, M.D., co-recipient of the 2005 Nobel Prize in Medicine, was online to the National Library of Medicine as early as the 1980s. He was a computer enthusiast and, by searching Medline from Australia, found—and connected—"widespread, dispersed references to things in the stomach, which seemed to be related to the bacteria…we could then develop a hypothesis that these bacteria were causing some problem in the stomach, and maybe that was leading to ulcers." Another recent account of the value of NLM's services was received in October 2005 when the medical librarian at the University of Pittsburgh Medical Center described how a patient at their trauma center faced losing an arm following an industrial accident in which the patient's arm was trapped in a meat grinder. A search of the PubMed/Medline database turned up a 1975 article in the Journal of Trauma, complete with a picture, that was credited with leading the Pittsburgh physicians to successfully extricate the arm without further damage to the remaining viable tissues.
The following sections describe the Library's many information services for both the general public and the professional/scientific community, the R&D that is carried out by NLM staff to create new and improved services, and the broad and varied support the Library provides to the academic and medical community to enhance medical communications.
STORIES OF DISCOVERY
The National Library of Medicine has long been an innovator in disseminating medical information—from the pioneering Index Medicus, begun in 1879 and relied on by the medical community for a century and a quarter, to the first successful application of computers to run a large-scale bibliographic system in the mid-20th century, to an unmatched array of Web-based medical information services in the 21st century. The "stories of discovery" that follow include databases and research tools for scientists and health professionals and a growing variety of Web information services intended for the general public.
For the Public: Good Health Information—from the NIH and Other Reliable Sources
MedlinePlus
MedlinePlus.gov is a Web-based service, available in both English and Spanish, that provides integrated access to authoritative consumer health information. Much of this material comes from the NIH Institutes and, although the information is based on the medical research the Institutes sponsor and carry out, it has been carefully written and formatted to be easily understood by the public. NLM relies on its cadre of experienced medical librarians to select and organize this information for the Web. In addition, they receive help from members of the National Network of Libraries of Medicine across the country. That they have been successful may be seen by the fact that MedlinePlus and MedlinePlus en español are the Federal Web sites most highly ranked by the independent American Customer Satisfaction Index.
In the seven years since its introduction, MedlinePlus has grown tremendously, both in terms of its coverage of health information and its usage by the public. In October 2005 there were 67 million page views by 8.6 million unique visitors. It has more than 700 "health topics," containing, for example, overview information, pertinent clinical trials, alternative medicine, prevention, management, therapies, the latest research, and the latest news from the print media. There are even links to the scientific literature through PubMed/Medline (described below). In addition to the health topics, there are medical dictionaries, a medical encyclopedia, directories of hospitals and providers, and interactive "tutorials" with images and sound. The newest addition to MedlinePlus is a series of surgical videos that show actual operations of common surgical procedures. MedlinePlus en español was introduced in 2002 and has grown to virtual parity with the English version. Another new aspect of MedlinePlus is "Go Local," that is, a service to link users from the MedlinePlus health topics to the health and social services in their community that are related to that topic. Ten states are represented in Go Local, and more are coming online all the time.
Although MedlinePlus is already one of the most heavily trafficked Web sites containing health information for the public, the Library actively seeks to promote the database. For example, the NLM is collaborating with the American College of Physicians Foundation in "Information Rx," a project to encourage physicians to make information referrals to MedlinePlus. Since patients trust their physicians to recommend good health information, the idea is to promote MedlinePlus as the "Web site your doctor prescribes." The Foundation sees this as an opportunity to help member physicians provide reliable health information for their patients. NLM provides a variety of information materials to the participating physicians, including posters and prescription pads. The project continues to be expanded and the results so far are encouraging—two thirds of the physicians ranked MedlinePlus as their first or second choice for referring patients. NLM is also working on a similar program in Florida with the members of the American Medical Association Foundation.
NIHSeniorHealth.gov
NIHSeniorHealth.gov, another Web site for consumers, is maintained by the National Library of Medicine in collaboration with the National Institute on Aging. At present there are 22 topics of interest to seniors, prepared in cooperation with 9 NIH Institutes, including, for example, Alzheimer's Disease, balance problems, macular degeneration, shingles, and stroke. More topics are in preparation. There are also videos of inspiring "exercise stories" from seniors as old as 91. NIHSeniorHealth.gov contains information in a format that is especially usable by seniors. For example, the site features large print and easy-to-read segments of information repeated in a variety of formats—such as open-captioned videos and short quizzes—to increase the likelihood it will be remembered. NIHSeniorHealth.gov also has a "talking" function, which allows users the option of reading the text or listening to it as it is read to them.
ClinicalTrials.gov
Launched in February 2000, ClinicalTrials.gov was created to give everyone easy access to information about human research studies. ClinicalTrials.gov has proven popular with the public, logging approximately 7 million page views/month by approximately 20,000 visitors daily. Without question, it has helped investigators to recruit patients, too. The site contains information on more than 23,500 federally and privately supported trials. It includes summaries of the purpose of each study, the recruiting status, criteria for patient participation, location(s) of the trial and specific contact information. ClinicalTrials.gov was originally implemented to fulfill legislative requirements that called for a registry of publicly and privately sponsored trials of investigational new drugs (IND) for serious or life-threatening diseases submitted to the FDA. However, ClinicalTrials.gov now serves broader functions as a trials registry. All trials, regardless of study design, sponsor, intervention type, or IND status, are accepted into the registry. In addition, sponsors are encouraged to submit links to peer-reviewed articles reporting results for completed trials. New features have been introduced to help users find studies in different ways, such as geographical area (Browse by Map) and patient recruitment status (Browse by Status).
In September 2004, the International Committee of Medical Journal Editors (ICMJE), whose members include editors from such prestigious publications as the New England Journal of Medicine, the Journal of the American Medical Association and The Lancet, announced a policy requiring trial registration as a condition of publication of the results of clinical trials. The response to the ICMJE requirements has been substantial: the number of registered trials at ClinicalTrials.gov increased by 81 percent from 13,000 to over 23,500 in five months (May to October 2005).
Genetics Home Reference
Findings from the Human Genome Project have greatly increased the public's interest in genetics, but the technical data resulting from the Project can be overwhelming to those without formal training. The Genetics Home Reference is a Web-based information system intended to help with this problem. It provides brief, consumer-friendly summaries of genetic conditions and related genes and chromosomes. This information resource bridges consumer health information (for example, MedlinePlus) and bioinformatics data (for example Entrez Gene), and it links to many existing resources, both at NLM and at other reliable sites. At its launch in February 2003, Genetics Home Reference covered a handful of diseases that had a prominent genetic component. As of October 2005 it covered about 450 topics grouped in categories, for example, cancers, brain and nervous system, lungs and breathing, and digestive system. Understandability is a primary consideration for information in Genetics Home Reference, and all topics receive close review by experts before they are put in the database. An online handbook to help consumers understand genetics has been created and made available in the process. The Genetics Home Reference is developed and managed by NLM in close cooperation with other Federal health agencies, among them the Health Resources and Services Administration, National Cancer Institute, the National Human Genome Research Institute, and the Office of Rare Diseases.
Toxicology and Environmental Health
The Library's Division of Specialized Information Services has introduced a number of Web-based information services for the general public. One of these is the colorful "Tox Town." Tox Town looks at an ordinary town and points out many harmful substances and environmental hazards that might exist there. Users can click on a town location, like the school, and see a colorful dollhouse-style cutaway view of that building. Toxic chemicals that might be found in the school are listed, along with links to selected Internet resources about school environments. There are similar cutaways for offices, factories, parks, and other locations. NLM has introduced several other versions of Tox Town that concentrate on the special problems found in a big city, the U.S.–Mexico border area, and a farm. There is also a new special section with information on toxic chemicals and disaster health concerns in the wake of Hurricane Katrina and Hurricane Rita. A related database, TOXMAP, is an interactive Web site that shows—on maps—the amount and location of certain toxic chemicals released into the environment in the United States. Unlike Tox Town, which is hypothetical, TOXMAP actually focuses on the geographic distribution of chemical releases, their relative amounts, and their trends over time. This release data comes from industrial facilities around the United States, as reported annually to the Environmental Protection Agency. TOXMAP is being used by public health professionals, public health students and faculty, librarians, community health nurses, and citizens who are concerned about the location of toxic chemicals in their neighborhood.
Another consumer information resource created by the Division of Specialized Information Services is the Household Products Database. This is a guide that provides easy-to-understand data on the potential health effects of more than 2,000 ingredients contained in more than 6,000 common household products. The database provides information on many of these substances and their potential health effects, in consumer-friendly language. For convenience in searching, the products are categorized—auto, inside the home, personal care, pet care, etc. The Household Products Database has proved to be popular with the media, and there have been a number of newspaper and magazine articles about it. For more technical information, users can launch a search for a product or ingredient from the product's page into NLM's TOXNET®, a cluster of databases with scientific information on toxicology, hazardous chemicals, and related information.
Profiles in Science
Profiles in Science® is a unique site that celebrates twentieth-century leaders in biomedical research and public health. It makes the archival collections of prominent scientists, physicians, and others who have advanced the scientific enterprise available to the public through modern digital technology. It promotes the use of the Internet for research and teaching in the history of biomedical science. Many of the collections have been donated to NLM and contain published and unpublished items, including books, journal volumes, pamphlets, diaries, letters, manuscripts, photographs, audiotapes, video clips, and other materials. Each digital collection on the Profiles site consists of two major parts. The first part is an exhibit composed of introductory narratives on the scientist's life and work and a selection of noteworthy documents (text, audiotapes, video clips and photographs). This exhibit is particularly designed for students and those with little background in science. The second part consists of additional documents from the scientist's papers, available through a search engine and in alphabetical and chronological "views." The Profiles in Science collections are particularly strong in the areas of cellular biology, genetics, and biochemistry, but also reflect issues in such areas as health and medical research policy, the application of computers in medicine, science education, and the search for extraterrestrial life. Among the sixteen notables included are profiles on Joshua Lederberg, Albert Szent-Györgyi, and Francis Crick.
Special Populations
One "special population," seniors, and the popular database, NIHSeniorHealth.gov, have already been discussed. Another subset of our population, those whose primary language is Spanish, has also been noted, with MedlinePlus en español. NLM has also created other Web-based information services for specialized populations. Among these are American Indian Health, Arctic Health, and Asian American Health. These sites contain extensive information about diseases and conditions for both the public and health professionals of special concern to those populations and links to organizations and government agencies that can provide assistance. One way these services differ is their emphasis on traditional healing and complementary medicine.
Information Services for Health Professionals and the Scientific Community
PubMed/Medline
In 1879 the Library began publishing the Index Medicus, a listing of medical journal articles. This was, at the time, an unprecedented undertaking, and came to be recognized as one of the major U.S. contributions to 19th century medicine. Almost a century later, the references to journal articles were computerized and made available through an online service called Medline. Medline was the first successful marriage of a large reference database with a national telecommunications network. Originally, trained librarians were required to search the database. However, in the eighties, NLM introduced "Grateful Med," a software program created by the NLM that one could load onto a PC and, equipped with a modem and a password, search Medline right from one's home, office, or laboratory. Grateful Med was eagerly snapped up not only by librarians but also by health professionals, scientists, students, lawyers, medical journalists and others, who saw the average charge of $2 per Medline search as a bargain.
In 1997, free Medline searching via the World Wide Web was introduced using a new system called PubMed®. PubMed is a sophisticated yet easy to use literature retrieval system developed by NLM's National Center for Biotechnology Information to provide access to citations and abstracts for biomedical science journal literature. For the first time, anyone with access to the Web could search freely through what is now a database of more than 15 million references and abstracts for medical journal articles from the 1950s to the present. Usage of PubMed/Medline by the scientific and lay communities has grown considerably since its introduction in 1997, to over 2.8 million searches per day.
PubMed also links to the sites of participating publishers so that users can retrieve full-text articles from 5,000 journals. Where links to electronic full text are not available, the user may use PubMed to place an online order for an article directly from a library in the National Network of Libraries of Medicine. PubMed allows users to customize their access, for example with an option to automatically update and e-mail search results from user-saved searches. A spell check feature was added to PubMed that provides the user with choices of alternate spellings for search terms. The ability of medical scientists and health professionals to have instant access to scientific articles (some free, some with a modest charge) is unparalleled and not matched by any other scientific discipline. The National Institutes of Health Web site is the second most heavily trafficked site in the Federal Government, and the NLM databases account for the major share of that use.
PubMed Central
PubMedCentral® (PMC) is a Web-based repository of biomedical journal literature providing free and unrestricted access to the full-text of articles. This repository is based on a natural integration with the existing PubMed biomedical literature database of references and abstracts. Currently, PMC contains over 500,000 full-text articles from 214 biomedical journals. Recent additions have come from newly published material as well as from digitizing back issues that previously were only available in printed form. Among the journals recently joining PMC was the first journal from the UK Wellcome Trust's collaborative effort to digitize the complete backfiles of a number of important and historically significant medical journals. In addition to creating a digital copy of every page in the backfiles, the digitization process will also create a PDF file for every discrete item (article, editorial, letter, advertisement, etc.) in the archive, and use optical character recognition technology to generate searchable text. NIH's Public Access policy to publications resulting from NIH-funded research went into effect on May 2, 2005. NCBI designed and implemented the NIH Manuscript Submission system and is responsible for archiving the manuscripts in the PubMedCentral database. The system provides a quick and easy-to-use system for manuscript submission. Creating such digital archives as PubMedCentral to ensure that the world's biomedical literature is properly recorded and available for future generations, is an important NLM responsibility.
GenBank
GenBank® is the NIH genetic sequence database, an annotated collection of all publicly available DNA sequences. NLM's National Center for Biotechnology Information is responsible for all phases of GenBank production, support, and distribution, including timely and accurate processing of sequence records and biological review of both new sequence entries and updates to existing entries. Integrated retrieval tools allow seamless searching of the sequence data housed in GenBank and provide links to related sequences, bibliographic citations, and other resources. Such features allow GenBank to serve as a critical research tool in the analysis and discovery of gene function as well as discoveries that lead to identification and cures for a number of diseases. One recent example (November 2005) of the use of NCBI sequence databases was in the identification of the first polio case in the U.S. since 1999. The state health laboratory in Minnesota had isolated an unknown virus from a hospitalized child from an Amish community. When the laboratory determined the virus's DNA code, they went to the Web, searched against the 55 million DNA sequences at NCBI and found a match to the polio virus used in the Sabin oral vaccine. "Bingo," said the laboratory's director, "It was a 98 percent match. We knew we had nailed it."
Important sources of data for GenBank are direct sequence submissions to NCBI from thousands of individual scientists and genome sequencing centers prior to publication. GenBank indexers with specialized training in molecular biology create the GenBank records and apply rigorous quality control procedures to the data. Records submitted to NCBI's international collaborators in the United Kingdom and Japan are shared through an automated system of daily updates. Other cooperative arrangements, such as those with the U.S. Patent and Trademark Office for sequences from issued patents, augment the data collection effort and ensure the comprehensiveness of the database. GenBank reached a total of 100 billion bases this year, a milestone for the database and its collaborators.
Resources to Support Genome Analysis
The National Center for Biotechnology Information has developed a suite of genomic resources to support comprehensive analysis of the human genome, as well as the complete genomes of several model organisms. Specialized tools and databases have also been designed to facilitate researchers' use of this data. NCBI maintains an expanding collection of specialized, yet integrated, database repositories that collectively capture and redistribute the biological relationships between genome sequences, expressed mRNAs and proteins, and individual sequence variations. Examples:
- NCBI's Web resource, "Human Genome Resources," serves as a nexus for the collection and storage of genomic human data. This site provides access to information on sequencing efforts and various other types of resources, such as chromosome-specific mapping information, cytogenetics, and genome similarity plotting.
- The Reference Sequence (RefSeq) database is a comprehensive, integrated, non-redundant set of sequences, including genomic DNA, gene transcript (RNA), and protein products for 3,000 major research organisms. These standard sequences serve as a basis for medical, functional, and diversity studies by providing a stable reference for gene identification and characterization, mutation analysis, expression studies, polymorphism discovery, and comparative analysis.
- Entrez Gene integrates information about 1.3 million genes and gene features annotated on RefSeq making it easier for researchers to find and interpret gene-specific information.
- The Map Viewer is a heavily used tool for genome research, enabling researchers to visualize the genome in its entirety as well as down to the DNA base level. Users can search for genes or gene markers by submitting a query against the entire genome of an organism or against specific regions. Results can be displayed graphically providing the sequence context for the match and linked data to other regions or to other organisms
- Genes and Disease is a collection of articles designed to educate the lay public and students on how genes are inherited and cause disease and how an understanding of the human genome will contribute to improving diagnosis and treatment of disease. This collection, part of the NCBI Books site, contains descriptions for over 150 genetic diseases and links to databases and organizations for additional information.
- OMIM is an electronic version of Dr. Victor McKusick's "Mendelian Inheritance in Man," a catalog of human genes and genetic disorders. The database, produced at the Johns Hopkins School of Medicine, contains over 15,500 detailed descriptions of genetic disorders as well as maps showing the cytogenetic location of disease genes.
- The GeneTests database produced at the University of Washington is now being supported by contract from NCBI. GeneTests is used more than 25,000 times a day by genetics counselors and physicians for its comprehensive genetic disease descriptions and for locating genetics clinics and testing laboratories.
Grant Support for Training
NLM remains the principal source of support nationally for research training in the field of biomedical informatics. This support is especially important as rapidly moving technology in health care and biomedical research requires investigators who understand biomedicine as well as fundamental problems of knowledge representation, decision support, and human-computer interface. Through the Library's Extramural Programs Division, five-year institutional training grants support pre-doctoral, post-doctoral, and short-term trainees in 18 programs across the country. In total, 27 institutions provide training with NLM funds, supporting almost 300 trainees. Collectively, the programs emphasize training in clinical informatics, bioinformatics and computational biology, and public health informatics. In a related program, the NLM and the Robert Wood Johnson Foundation have recently formed a partnership to support training in public health informatics. Through a $3.6 million grant from the Foundation, four existing NLM-funded training sites received supplemental awards to develop formal training tracks in public health informatics and for support of trainees in these tracks.
SCIENCE ADVANCES
PubChem: A Milestone on the NIH Roadmap
A critical need in biomedical research, as identified in the NIH Roadmap Initiative, is a repository for what are called "small molecules," which are crucial in drug development. Small molecules are responsible for the most basic chemical processes that are essential for life, such as those that produce energy for the cell and those that carry messages to cells and their components. Considering their basic role in the cell, it comes as no surprise that small molecules often play an essential role in the attack of a pathogen, or in the cell's response to the attack. Most therapeutic drugs are small molecules that interact with macromolecules and influence their behavior. It is therefore of great importance to link small molecules and their chemistry to the structures of large macromolecules, such as the protein and DNA structures found in NCBI's biological databases and in the scientific literature.
The new PubChem does this. It is a key component of the NIH Roadmap project in Molecular Libraries and Imaging. PubChem links the small molecules to their biological functions and to the macromolecules with which they interact. The PubChem suite—Substance, Compound, and BioAssay—contain substance information, small molecule structures, and bioactivity data, such as the results of high-throughput biological screening experiments. The PubChem suite of databases is part of Entrez, NCBI's search and retrieval system, which provides compound neighboring, and allows searches by structural similarity or bioactivity. Structures stored within PubChem Compound are clustered and cross-referenced by identity and similarity groups. Additionally, calculated properties and descriptors are available for searching. PubChem's chemical structure records are linked to other NLM databases such as the PubMed database of scientific literature, and NLM's protein 3D structure Molecular Modeling Database. At present, PubChem includes over 7.5 million records for small molecules with over 5 million molecular structures. PubChem has built its database of small molecule structures, properties, and activities from data in public, academic, and commercial resources (including NLM's ChemIDplus, the National Cancer Institute's Developmental Therapeutics Program, and the National Institute of Standards and Technology's Mass Spectral Library). Data from NIH-funded small-molecule screening centers will add key information on biological properties.
A New Information Tool to Combat the Influenza Virus
More than 30,000 Americans die each year of flu-associated disease (JAMA 289:179-186, 2003. In the past 100 years there have been three major pandemics which affected between 20% and 40% of the world's population in a single year. The Spanish Flu pandemic of 1918–1919, for example, affected over 200 million people, killing more than 50 million worldwide (Bull. Hist. Med. 76: 105-115, 2002. Notable pandemics also occurred in 1957 and 1968. A detailed understanding of the causes of influenza epidemics and pandemics is crucial to their control. The development of vaccines for influenza, however, presents a problem. Because the virus changes from year to year, vaccines that target last year's virus are usually ineffective against this year's variants. Recently, advances in molecular biology, genomic sequencing, and genomic analysis have begun to provide insights into the processes leading to rapid changes in the influenza virus.
As a foundation for such studies, the National Institute of Allergy and Infectious Diseases (NIAID), in collaboration with the NCBI and several other partners, has funded the Influenza Genome Sequencing Project which aims to deposit a large number of influenza sequences into NCBI's GenBank sequence database. To date, the project has generated over 3000 influenza sequences including those for more than 150 complete influenza genomes. To help researchers make the best use of this data, NCBI's Influenza Virus Resource links the NIAID project data to the most recent scientific literature on influenza as well as to a number of online analysis tools and databases, including influenza sequences in GenBank and RefSeq.
Research using the tools of genomic analysis can lead to the discovery of important mechanisms of viral change. Initial work reveals the process of influenza viral evolution to be complex, and further progress depends on the collection of more extensive datasets and the further integration of new sequence data with other biological information. As the data accumulates and the analyses progress, the discoveries made will ultimately lead to better prediction of large-scale outbreaks, more effective vaccine design, and the saving of many human lives.
A New Tool for the Rapid, Reliable Analysis of DNA
The NCBI and the National Institute of Justice (NIJ) have worked together to develop public domain software to make forensic analyses of DNA profiles rapid and reliable. The need for such software was evident in case of the thousands of victims of the World Trade Center attacks. The need, however, is broader than just the identification of victims of disasters. It extends to the need for independent review of DNA profiles for convicted felons in state DNA profile databases. The problem is a challenging one because of the variation in the equipment and forensic procedures employed by regional laboratories. Effective software, in addition to processing the data rapidly, must be robust enough to adapt to the changing biases in the forensic data resulting from these variable laboratory environments.
To meet this need, scientists at NCBI and NIJ have developed a software package called the Open Source Independent Review and Interpretation System (OSIRIS). When used in conjunction with a forensic laboratory's data collection software, OSIRIS rapidly assesses the accuracy of forensic analyses and measures the conformance of the raw data to quality control standards, flagging problems in the data and automatically highlighting informative data features. OSIRIS then enables both the data and its interpretation to be exported in a structured manner to facilitate latter reanalysis and to provide shared access between authorized parties. Shared access to the data allows quality management by state labs and regulatory agencies, reduces caseloads through automated review, and leads to increased throughput without compromising accuracy. OSIRIS is rapid—it can handle about 10,000 analyses in 2½ minutes while it typically takes a technician 20 minutes to do a single analysis. The software is now being tested within five collaborating forensic DNA laboratories, and it is being used to assist in the analysis and validation of forensic data from the Gulf Coast states in the aftermath of Katrina.
OTHER AREAS OF INTEREST
NLM and Disaster Management
NLM has actually been involved in disaster response and management since the late 1960s, when the Library's Specialized Information Services Division was established as the U.S. government focal point for information on toxic substances and environmental health. The Library's services in this area have been used in such emergencies as the gas leak disaster in Bhopal, India, in the eighties, and Hurricane Mitch and the earthquakes in Central America in the nineties. In the aftermath of September 11, 2001, NLM increased the focus on use of information services and technology to enhance disaster response and management (defined to encompass all types of natural and manmade disasters) in its extramural and intramural activities. Some of these activities include:
- The development of OSIRIS to identify victims' remains via DNA, described above.
- Four grant programs to support information management during natural disasters or terrorist attacks, including one explicitly named Informatics for Disaster Management Grants.
- WISER (Wireless Information System for Emergency Responders) provides information via mobile devices to help identify unknown harmful substances to those responding to chemical emergencies. WISER was used in Louisiana after Hurricane Katrina.
- NLM itself has taken steps to protect its databases and information services from outside attack or interference, including establishing an extensive off-site backup facility.
Standards for Electronic Health Records
As part of its Unified Medical Language System® (UMLS) project, the National Library of Medicine produces and distributes vocabulary databases and software tools designed to assist informatics researchers and system developers in automated interpretation and integration of medical knowledge and health data. Chief among the UMLS resources is the Metathesaurus, which links and provides 4.7 million concept names for 1.2 million concepts from 114 vocabularies in a single database format. The UMLS serves as a common distribution vehicle for standard code sets and vocabularies needed for administrative transactions and electronic health records, as well as a resource for advanced natural language processing, automated indexing, and enhanced information retrieval.
Building on two decades of UMLS experience, the National Library of Medicine serves as an HHS coordinating center for standard clinical vocabularies. NLM licenses SNOMED CT® clinical terminology for US-wide use; funds the ongoing development of LOINC standard identifiers, names, and codes for clinical tests and results; builds and maintains the RxNorm clinical drug vocabulary. The Library works closely with the Office of the National Coordinator for Health Information Technology and relevant federal agencies and organizations to align health data standards into an effective interlocking set and to promote more rapid adoption of standards-based electronic health records to facilitate patient care, public health surveillance, and clinical research.
In FY 2005, NLM served as the operational home for the Commission on Systemic Interoperability, established by the Medicare Modernization Act of 2004 to develop an overall strategy for achieving interoperable electronic health records. The Commission, with members appointed by the President and leaders in the House and Senate, delivered its final report in October 2005. Its recommendations will inform the work of the American Health Information Community, recently established by the Secretary of Health and Human Services to advise on specific priority action steps to achieve interoperable, digital health records.
Long Range Planning
The National Library of Medicine, 20 years ago, published a long range plan that has proved to be of enormous benefit to the institution. Out of it grew such initiatives as the Visible Human Project, the National Center for Biotechnology Information, and the recommendation that the Library engage in an outreach campaign to reach minority and other underserved health professionals. The Library is now engaged in a similar planning exercise for the next quarter century. Leaders from across the spectrum of health and medicine are meeting at the Library to consider four major themes: NLM resources and infrastructure for the 21st century, NLM outreach to the underserved in the 21st century, NLM support for clinical and public health systems for the 21st century, and NLM support for genomics in the 21st century. The plan will be reviewed by the NLM Board of Regents and published in 2006.
NLM BUDGET POLICY
The Fiscal Year 2007 budget request for the NLM is $313,269,000, a decrease of $1,641,000 and .5 percent less than the FY 2006 Appropriation. Included in the FY 2007 request is NLM's support for the trans-NIH Roadmap initiatives, estimated at 1.2% of the FY 2007 budget request. A full description of this trans-NIH program may be found in the NIH Overview.
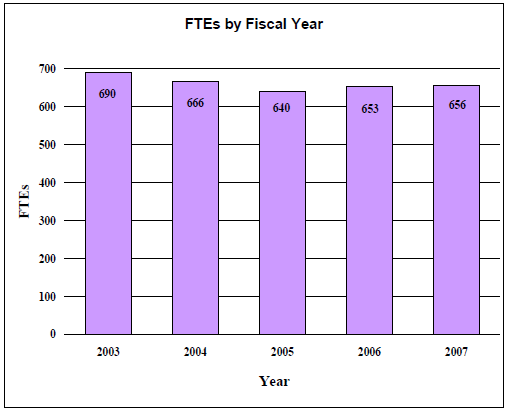
A five year history of FTEs and Funding Levels for NLM are shown in the graphs below. Note that as the result of several administrative restructurings in recent years, FTE data is non comparable.
Data for FTEs by Fiscal Year Chart
| Year | FTEs |
|---|---|
| 2003 | 690 |
| 2004 | 666 |
| 2005 | 640 |
| 2006 | 653 |
| 2007 | 656 |
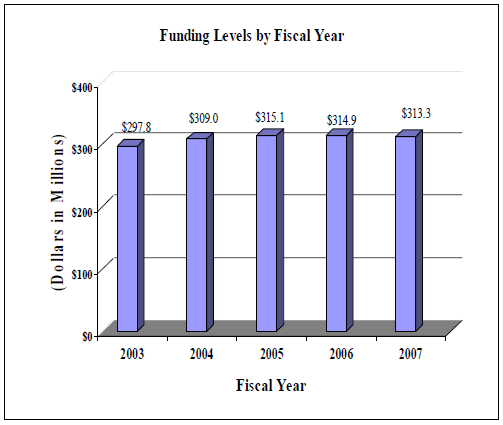
Data for Funding Levels by Fiscal Year Chart (Dollars in Millions)
| Year | FTEs |
|---|---|
| 2003 | $297.8 |
| 2004 | $309.0 |
| 2005 | $315.1 |
| 2006 | $314.9 |
| 2007 | $313.3 |
The request continues to support health care applications; improve consumer health initiatives; utilize advanced computer and communications technologies; emphasize support for library operations; funds grants in training, medical informatics, biotechnology and other research developments activities.
NIH must nurture a vibrant, creative research workforce, including sufficient numbers of new investigators with new ideas and new skills. In the FY 2007 budget request for NLM, $450,000 will be used to support 5 awards for the new K/R "Pathway to Independence" program.
NLM will also support the Genes, Environment, and Health Initiative (GEHI) to: 1) accelerate discovery of the major genetic factors associated with diseases that have a substantial public health impact; and 2) accelerate the development of innovative technologies and tools to measure dietary intake, physical activity, and environmental exposures, and to determine an individual's biological response to those influences. The FY 2007 request includes $530,000 to support this project.
The FY 2007 budget request for NLM by budget activity is illustrated below.
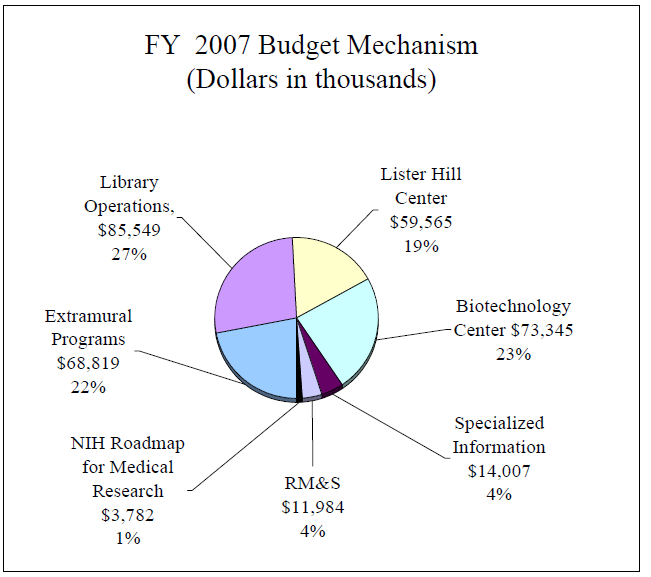
Data for FY 2007 Budget Mechanism (Dollars in Thousands) Chart
| Programs | Amount (Dollars in Thousands) |
Percentage |
|---|---|---|
| Library Operations | $85,549 | 27% |
| Extramural Programs | $68,819 | 22% |
| Lister Hill Center | $59,565 | 19% |
| Biotechnology Center | $73,345 | 23% |
| Specialized Information | $14,007 | 4% |
| RM&S | $11,984 | 4% |
| NIH Roadmap for Medical Research | $3,782 | 1% |
Budget Mechanism - Total
| Mechanism | FY 2005 Actual |
FY 2006 Appropriation |
FY 2007 Estimate |
|||||
|---|---|---|---|---|---|---|---|---|
| No. | Amount | No. | Amount | No. | Amount | |||
| Grants: | ||||||||
| Noncompeting | 146 | $41,333,000 | 132 | $40,068,000 | 125 | $40,038,000 | ||
| Competing | 61 | 11,207,000 | 48 | 10,833,000 | 47 | 10,693,000 | ||
| SBIR/STTR | 4 | 728,000 | 6 | 647,000 | 5 | 600,000 | ||
| Subtotal, Grants | 211 | 53,268,000 | 186 | 51,548,000 | 177 | 51,331,000 | ||
| Contracts: | ||||||||
| Noncompeting | 8 | 51,331,000 | 0 | 0 | 8 | 12,541,000 | ||
| Competing | 8 | 3,690,000 | 15 | 17,699,000 | 7 | 4,947,000 | ||
| Subtotal, Contracts | 16 | 16,132,000 | 15 | 17,699,000 | 15 | 17,488,000 | ||
| Total, Extramural | 227 | 69,400,000 | 201 | 69,247,000 | 192 | 68,819,000 | ||
| FTEs | FTEs | FTEs | ||||||
| Intramural programs | ||||||||
| Library Operations | 330 | 84,934,000 | 327 | 85,594,000 | 329 | 85,549,000 | ||
| Lister Hill Center | 72 | 61,106,000 | 69 | 57,511,000 | 70 | 55,783,000 | ||
| Biotechnology Center | 148 | 72,273,000 | 150 | 73,751,000 | 150 | 73,345,000 | ||
| Specialized Information | 31 | 14,550,000 | 33 | 14,186,000 | 33 | 14,007,000 | ||
| Total, Intramural | 581 | 232,863,000 | 579 | 231,042,000 | 582 | 228,684,000 | ||
| Research management and support | 80 | 10,891,000 | 74 | 11,807,000 | 74 | 3,782,000 | ||
| NIH Roadmap for Medical Research | 1,992,000 | 2,814,000 | 3,782,000 | |||||
| Total, NLM | 661 | 315,146,000 | 653 | 314,910,000 | 656 | 313,269,000 | ||
Includes FTEs which are reimbursed from the NIH Roadmap for Medical ResearchAppropriation
Budget Authority by Activity
| Activity (Dollars in Thousands) |
FY 2005 Actual |
FY 2006 Appropriation |
FY 2007 Estimate |
Change | ||||
|---|---|---|---|---|---|---|---|---|
| FTEs | Amount | FTEs | Amount | FTEs | Amount | FTEs | Amount | |
| Extramural Research | ||||||||
| Medical Library Assistance | $39,247 | $38,514 | $38,029 | (485) | ||||
| PHS 301 | 29,425 | 30,086 | 30,190 | 104 | ||||
| SBIR/STTR | 728 | 647 | 600 | (47) | ||||
| Subtotal, Extramural | 69,400 | 69,247 | 68,819 | (428) | ||||
| Intramural Programs: | ||||||||
| Library Operations | 330 | 84,934 | 327 | 85,594 | 329 | 85,549 | 2 | (45) |
| Lister Hill Center | 72 | 61,106 | 69 | 57,511 | 70 | 55,783 | 1 | (1,728) |
| Biotechnology Center | 148 | 72,273 | 150 | 73,751 | 150 | 73,345 | 0 | (406) |
| Specialized Information | 31 | 14,550 | 33 | 14,186 | 33 | 14,007 | 0 | (179) |
| Subtotal, Intramural | 581 | 232,863 | 579 | 231,042 | 582 | 28,684 | 3 | (2,358) |
| Research management & support | 80 | 10,891 | 74 | 11,807 | 74 | 11,984 | 0 | 177 |
| NIH Roadmap for Medical Research | 1,992 | 2,814 | 3,782 | 968 | ||||
| TOTAL | 661 | 315,146 | 653 | 314,910 | 656 | 313,269 | 3 | (1,641) |
Includes FTEs which are reimbursed from the NIH Roadmap for Medical Research
Summary of Changes
FY 2006 Appropriation: $314,910,000
FY 2007 Estimated Budget Authority: 313,269,000
Net change: (1,641,000)
| CHANGES | FY 2006 Appropriation |
Change from Base | ||
|---|---|---|---|---|
|
FTEs |
Budget Authority |
FTEs | Budget Authority |
|
| A. Built-in: | ||||
| 1. Intramural programs: | ||||
| a. Within grade increase | $61,275,000 | $1,073,000 | ||
| b. Annualization of January 2006 pay increase | 61,275,000 | 483,000 | ||
| c. January 2007 pay increase | 61,275,000 | 1,037,000 | ||
| d. Payment for centrally furnished services | 10,069,000 | 151,000 | ||
| e. Increased cost of laboratory supplies, materials, and other expenses | 162,512,000 | 4,782,000 | ||
| Subtotal | 7,526,000 | |||
| 2. Research management and support: | ||||
| a. Within grade increase | 7,989,000 | 140,000 | ||
| b. Annualization of January 2006 pay increase | 7,989,000 | 63,000 | ||
| c. January 2007 pay increase | 7,989,000 | 135,000 | ||
| d. Payment for centrally furnished services | 0 | 0 | ||
| e. Increased cost of laboratory supplies, materials, and other expenses | 3,878,000 | 84,000 | ||
| Subtotal | 422,000 | |||
| Subtotal, Built-in | 7,948,000 | |||
| B. Program: | ||||
| 1. Research project grants: | ||||
| a. Noncompeting | 132 | $40,068,000 | (7) | ($30,000) |
| b. Competing | 48 | 10,833,000 | (1) | (140,000) |
| c. SBIR/STTR | 6 | 647,000 | (1) | (47,000) |
| Total | 186 | 51,548,000 | (9) | (217,000) |
| 2. Research and development contracts | 15 | 17,699,000 | 0 | (211,000) |
| Subtotal, extramural | (428,000) | |||
| FTEs | FTEs | |||
| 3. Intramural research | 572 | 231,042,000 | 3 | (9,884,000) |
| 4. Research management and support | 74 | 11,807,000 | 0 | (245,000) |
| 5. NIH Roadmap for Medical Research | 7 | 2,814,000 | 0 | 968,000 |
| Subtotal, program | 314,910,000 | (9,589,000) | ||
| Total changes | 653 | 3 | (1,641,000) | |
Budget Authority by Object
| FY 2006 Appropriation | FY 2007 Estimate |
Increase or Decrease | ||
|---|---|---|---|---|
| Total compensable workyears: | ||||
| Full-time employment | 653 | 656 | 3 | |
| Full-time equivalent of overtime and holiday hours | 3 | 3 | 0 | |
| Average ES salary | $151,107 | $155,792 | $4,684 | |
| Average GM/GS grade | 10.9 | 10.9 | 0.0 | |
| Average GM/GS salary | $75,783 | $78,133 | $2,349 | |
| Average salary, grade established by act of July 1, 1944 (42 U.S.C. 207) | $93,432 | $96,328 | $2,896 | |
| Average salary of ungraded positions | 108,3811 | 11,741 | 3,360 | |
| OBJECT CLASSES |
FY 2006 |
FY 2007 Estimate |
Increase or Decrease |
|
| Personnel Compensation: | ||||
| 11.1 | Full-time permanent | $36,072,000 | $37,746,000 | $1,674,000 |
| 11.3 | Other than full-time permanent | 16,469,000 | 17,158,000 | 689,000 |
| 11.5 | Other personnel compensation | 1,673,000 | 1,745,000 | 72,000 |
| 11.7 | Military personnel | 191,000 | 199,000 | 8,000 |
| 11.8 | Special personnel services payments | 1,850,000 | 2,120,000 | 270,000 |
| Total, Personnel Compensation | 56,255,000 | 58,968,000 | 2,713,000 | |
| 12.0 | Personnel benefits | 12,936,000 | 13,526,000 | 590,000 |
| 12.2 | Military personnel benefits | 73,000 | 76,000 | 3,000 |
| Subtotal, Pay Costs | 69,264,000 | 72,570,000 | 3,306,000 | |
| 21.0 | Travel and transportation of persons | 1,241,000 | 1,229,000 | (12,000) |
| 22.0 | Transportation of things | 171,000 | 169,000 | (2,000) |
| 23.1 | Rental payments to GSA | 4,000 | 4,000 | 0 |
| 23.2 | Rental payments to others | 102,000 | 101,000 | (1,000) |
| 23.3 | Communications, utilities and miscellaneous charges | 1,966,000 | 1,946,000 | (20,000) |
| 24.0 | Printing and reproduction | 414,000 | 410,000 | (4,000) |
| 25.1 | Consulting services | 44,754,000 | 44,306,000 | (448,000) |
| 25.2 | Other services | 44,147,000 | 39,921,000 | (4,226,000) |
| 25.3 | Purchase of goods and services from government accounts | 46,206,000 | 45,744,000 | (462,000) |
| 25.4 | Operation and maintenance of facilities | 1,571,000 | 1,555,000 | (16,000) |
| 25.5 | Research and development contracts | 20,050,000 | 19,849,000 | (201,000) |
| 25.7 | Operation and maintenance of equipment | 10,724,000 | 10,617,000 | (107,000) |
| 25.0 | Subtotal, Other Contractual Services | 10,724,000 | 10,617,000 | (107,000) |
| 26.0 | Supplies and materials | 1,883,000 | 1,864,000 | (19,000) |
| 31.0 | Equipment | 18,040,000 | 17,860,000 | (180,000) |
| 32.0 | Land and structures | 0 | 0 | 0 |
| 41.0 | Grants, subsidies and contributions | 51,548,000 | 51,331,000 | (217,000) |
| 43.0 | Interest and dividends | 11,000 | 11,000 | 0 |
| Subtotal, Non-Pay Costs | 242,832,000 | 236,917,000 | (5,915,000) | |
| NIH Roadmap for Medical Research | 2,814,000 | 3,782,000 | 968,000 | |
| Total Budget Authority by Object | 314,910,000 | 313,269,000 | (1,641,000) | |
Includes FTEs which are reimbursed from the NIH Roadmap for Medical Research
Salaries and Expenses
| OBJECT CLASSES | FY 2006 Appropriation |
FY 2007 |
Increase or Decrease |
|---|---|---|---|
| Personnel Compensation: | |||
| Full-time permanent (11.1) | $36,072,000 | $37,746,000 | $1,674,000 |
| Other than full-time permanent (11.3) | 16,469,000 | 17,158,000 | 689,000 |
| Other personnel compensation (11.5) | 1,673,000 | 1,745,000 | 72,000 |
| Military personnel (11.7) | 191,000 | 199,000 | 8,000 |
| Special personnel services payments (11.8) | 1,850,000 | 2,120,000 | 270,000 |
| Total Personnel Compensation (11.9) | 56,255,000 | 58,968,000 | 2,713,000 |
| Civilian personnel benefits (12.1) | 12,936,000 | 13,526,000 | 590,000 |
| Military personnel benefits (12.2) | 73,000 | 76,000 | |
| Benefits to former personnel (13.0) | 0 | 0 | 0 |
| Subtotal, Pay Costs | 69,264,000 | 72,570,000 | 3,306,000 |
| Travel (21.0) | 1,241,000 | 1,229,000 | (12,000) |
| Transportation of things (22.0) | 171,000 | 169,000 | (2,000) |
| Rental payments to others (23.2) | 102,000 | 101,000 | (1,000) |
| Communications, utilities and miscellaneous charges (23.3) | 1,966,000 | 1,946,000 | (20,000) |
| Printing and reproduction (24.0) | 414,000 | 410,000 | (4,000) |
| Other Contractual Services: | |||
| Advisory and assistance services (25.1) | 44,754,000 | 44,306,000 | (448,000) |
| Other services (25.2) | 44,147,000 | 39,921,000 | (4,226,000) |
| Purchases from government accounts (25.3) | 28,131,000 | 27,458,000 | (673,000) |
| Operation and maintenance of facilities (25.4) | 1,571,000 | 1,555,000 | (16,000) |
| Operation and maintenance of equipment (25.7) | 10,724,000 | 10,617,000 | (107,000) |
| Subsistence and support of persons (25.8) | 0 | 0 | 0 |
| Subtotal Other Contractual Services | 129,327,000 | 123,857,000 | (5,470,000) |
| Supplies and materials (26.0) | 1,883,000 | 1,864,000 | (19,000) |
| Subtotal, Non-Pay Costs | 1,883,000 | 1,864,000 | (19,000) |
| Total, Administrative Costs | 204,368,000 | 202,146,000 | (2,222,000) |
Significant Items in House and Senate Appropriations Committee Reports
FY 2006 Senate Appropriations Committee Report Language (S. Rpt. 109-103)
Item
Disease Management Technology -- The Committee urges NLM to conduct outreach activities to all public and private sector organizations which have demonstrated capabilities in health information technology. The Committee is particularly interested in disease management technology as it relates to saving health care dollars, and improving care for chronically ill individuals and the workforce. (p. 159)
Action taken or to be taken
NLM regularly demonstrates its products and services at meetings, such as the Annual Symposium of the American Medical Informatics Association, that are aimed at public and private sector organizations with demonstrated capabilities in health information technology. The Library also offers training courses in the use of its Unified Medical Language System (UMLS) resources, which are designed for health information system developers. Some of NLM's services may assist developers of disease management technology in building and improving their products. In Fiscal Year 2006, the Library will work through relevant associations, such as the Disease Management Association of America, to highlight relevant NLM products to producers of disease management technology.
Item
Health Equity Review -- The Committee congratulates NLM for its concern about health disparities and its sensitivity to native populations, such as Native Americans, Alaskans, and Hawaiians, and other minority communities. The Committee encourages NLM to fund projects to map health disparities and population profiles in minority communities. (p. 159)
Action taken or to be taken
The Health Equity Review (HER) initiative started in 2004 as a collaboration of the Center for Public Service Communications, the National Library of Medicine, the National Indian Project Center (NIPC), Data Analysis Service (DAS), Bird's Eye View (BEV) and the native Hawaiian health organization Papa Ola Lokahi. More recently the University of Hawaii Department of Native Hawaiian Health joined the partnership. The goal of the initiative is to develop an electron HER Atlas that will display the specific locations where significant health disparaties exist. The base system would incorporate readily available national and state health data into an easy-to-use Geographic Information System (GIS) system. This system would then be made available, with appropriate training in its management and use, to minority communities which would add additional layers of community-derived health data, thereby allowing those communities to identify and portray their own health conditions and the resources available and needed to address them.
The current application project focuses on adding Native Hawaiian health, economic and traditional geographic boundaries to the existing HER Atlas. Much of this data is available only through local sources. In addition to developing the Atlas, NLM is supporting the training of a Native Hawaiian public health worker so that the community can take advantage of this tool for health planning and development. In addition this will aid in the sustainability of the program.
Future plans include supporting, linking to or otherwise promoting integration with the NLM TOXMAP program and its hazardous waste mapping data. NLM will identify additional communities to work with to test and refine the implementation and management of the tool as well as determine the level of training required for local implementation and management. If successful, the tool can be made freely available to other communities who wish to add and analyze their own data.
Item
Native Hawaiian Healing -- The Committee applauds the NLM's leadership for conducting ‘listening circles' to discuss the need for preservation and documentation of traditional cultural healing practices. The Committee encourages the NLM to explore the best ways of capturing and documenting this information through continued collaboration with Native Hawaiians. (p. 159)
Action taken to be taken
An NLM Working Group on the Library's role in documenting traditional Native American healing practices - including those of Native Hawaiians - has recommended changes to the library's collection development policy and practices such as the need to collect materials in additional formats and subject areas including ethnobotony, anthropology and religion. Changes are also required to the preservation policy to include these areas in long-term preservation programs. The manuscript collection policy already includes language that is inclusive of Native American traditional health and healing, although this has been refined and detailed. These collection development changes are currently being implemented. Further, NLM plans to support the development of the capacity of Native archives and libraries to develop and maintain their own archives and collections, with the understanding that the material should also be made widely available, possibly in digital format. In addition to developing its own collections, the Library will work with other interested agencies and organizations to plan national programs and develop standards. As an example, NLM is supporting the development of "Native American Protocols for American Libraries, Museums, and Information Services." Whenever possible, NLM will promote the digitization of information and materials and will link to such materials through its American Indian Health web site. To highlight traditional Native health and healing, in 2010 NLM will be releasing an exhibition on this subject. Input for and participation in this exhibition will be sought from U.S. Native communities and tribes.
Item
Outreach -- The Committee encourages NLM to continue its outreach activities aimed at educating health care professionals and the general public about the Library's products and services, in coordination with medical librarians and other health information specialists. (p.159)
Action taken or to be taken
The medical librarians and other health information specialists in the more than 5,000 libraries in the National Network of Libraries of Medicine (NN/LM) continue to be key players in NLM's efforts to educate health professionals and the general public about the Library's products and services. The basic goals of the NN/LM are to: (1) develop collaborations among network members and other organizations to improve access to biomedical information; (2) promote awareness of, access to, and use of biomedical information resources for health professionals and the public, with a particular emphasis on contributing to the Healthy People 2010 goal of eliminating health disparities; and (3) develop, promote and improve electronic access to health information by network members, health professionals, and organizations providing health in formation to the public.
In Fiscal Year 2005, NN/LM members received NLM funding for 132 special outreach projects in 28 states, the District of Columbia, the Virgin Islands, and Puerto Rico. In addition, NN/LM librarians demonstrated NLM products and services at more than 228 national, regional, state and local professional and consumer health meetings across the country.
In Fiscal Year 2006, new 5-year contracts will be awarded to 8 health sciences libraries to serve as Regional Medical Libraries in the National Network of Libraries of Medicine (NN/LM). In their first year, the new contracts are expected to award a minimum of 50-60 projects to academic health sciences, hospital, public and state libraries, public health departments, and community based organizations to improve electronic access to health information. The projects will focus on priority initiatives related to health disparities, health information literacy, HIV/AIDS, and public health and will cover an equal number of states. Special emphasis will be on projects that target minority and underserved populations. Training on NLM's information resources and support of computers and Internet connections will be major components of the projects.
Item
PubChem -- The Committee is aware of the development of PubChem, the informatics component of the Molecular Libraries project of the NIH Roadmap for Medical Research. The Committee understands that the purpose of PubChem is to create a database of chemical structures and their biological activities. PubChem will house both data from the new NIH molecular libraries screening center network and compound information from the scientific literature. The Committee expects the NIH to work with private sector chemical information providers, with a primary goal of maximizing progress in science while avoiding unnecessary duplication and competition with private sector databases. (p.159-160)
Action taken or to be taken
PubChem is a critical linchpin of the NIH Molecular Libraries Roadmap Initiative and is important to the progress of chemical genomics. PubChem facilitates NIH's goal to rapidly translate the discoveries of the genome into new therapeutics and to integrate small molecule chemistry into biomedical research. Its primary purpose is to organize new bioassay data generated by NIH-funded high throughput screening centers and connect these data to existing public genomic databases built and maintained by NLM's National Center for Biotechnology Information. Multiple private sector chemical information providers are currently contributing data to PubChem. There are also links from PubChem to private sector information resources.
Top leadership at the NIH has already consulted directly with several private sector chemical database providers regarding PubChem, including several interactions with the American Chemical Society (ACS). Given major differences in the scientific content of the two databases, NIH believes that PubChem is a complementary resource to the ACS Chemical Abstracts Database.
NIH also plans to involve private sector chemical information providers in the further development of PubChem. On September 1, 2005 the NIH issued a notice in the Federal Register requesting nominations for a special working group of the Board of Scientific Counselors of NLM's National Center for Biotechnology Information with the specific purpose "to advise on interactions with private sector information providers in the development of PubChem." The working group, which will have ACS representation, will advise on issues such as improving connections with the private sector chemical information providers to enhance linkages and interoperability among resources while avoiding any unnecessary duplication with commercial information services. The first meeting of the working group took place in December 2005.
Authorizing Legislation
| PHS Act/ Other Citation |
U.S. Code Citation |
2006 Amount Authorized |
FY 2006 Appropriation |
2007 Amount Authorized |
FY 2007 Budget Estimate |
|
|---|---|---|---|---|---|---|
| Research and Investigation | Section 301 | 42§241 | Indefinite | $314,910,000 | Indefinite | $313,269,000 |
| National Library of Medicine | Section 41(B) | 42§285b | Indefinite | Indefinite | ||
| National Research Service Awards | Section 487(d) | 42§288 | a/ | 0 | 0 | |
| Total, Budget Authority | $314,910,000 | $313,269,000 | ||||
a/ Amounts authorized by Section 301 and Title IV of the Public Health Act.
Appropriations History
| Fiscal Year | Budget Estimate to Congress |
House Allowance |
Senate Allowance |
Appropriation 1/ |
|---|---|---|---|---|
| 1998 | 152,689,000 2/ | 161,171,000 | 159,411,0002/ | 161,185,000 |
| 1999 | 170,738,000 | 176,492,000 | 181,309,000 | 180,742,000 |
| Rescission | (120,000) | |||
| 2000 | 185,654,0002/ | 202,027,000 | 210,183,000 | 215,214,000 |
| Rescission | (1,146,000) | |||
| 2001 | 230,135,000 2/ | 256,281,000 | 256,953,000 | 246,801,000 |
| Rescission | (399,000) | |||
| 2002 | 275,725,000 | 273,610,000 | 281,584,000 | 277,658,000 |
| Rescission | (1,567,000) | |||
| 2003 | 313,534,000 | 313,534,000 | 331,443,000 | 302,099,000 |
| Rescission | (1,964,000) | |||
| 2004 | 315,401,000 | 315,401,000 | 319,396,000 | 311,635,000 |
| Rescission | (2,520,000) | |||
| 2005 | 316,947,000 | 316,947,000 | 316,900,000 | 317,947,000 |
| Rescission | (2,801,000) | |||
| 2006 | 318,091,000 | 318,091,000 | 327,247,000 | 318,091,000 |
| Rescission | (3,181,000) | |||
| 2007 | 313,269,000 |
1/ Reflects enacted supplementals, rescissions, and reappropriations.
2/ Excludes funds for HIV/AIDS research activities consolidated in the NIH Office of AIDS Research
Details of Full-Time Equivalent Employment (FTEs)
| OFFICE/DIVISION | FY 2005 Actual |
FY 2006 Appropriation |
FY 2007 Estimate |
|---|---|---|---|
| Division of Library Operations | 330 | 327 | 329 |
| Lister Hill National Center for Biomedical Communications | 72 | 69 | 70 |
| National Center for Biotechnology Information | 148 | 150 | 150 |
| Division of Specialized Information Services | 31 | 33 | 33 |
| Office of the Director/Administration | 66 | 62 | 62 |
| Division of Extramural Programs | 14 | 12 | 12 |
| Total | 661 | 653 | 656 |
| Includes FTEs which are reimbursed from the NIH Roadmap for Medical Research | |||
| FTEs supported by funds from Cooperative Research and Development Agreements | (0) | (0) | (0) |
| FISCAL YEAR | Average GM/GS Grade | ||
| 2003 | 10.5 | ||
| 2004 | 10.6 | ||
| 2005 | 10.9 | ||
| 2006 | 10.9 | ||
| 2007 | 10.9 | ||
Details of Positions
| GRADE | FY 2005 Actual |
FY 2006 Appropriation |
FY 2007 Estimate |
|---|---|---|---|
| Total, ES Positions | 5 | 5 | 5 |
| Total, ES Salary | $730,409 | 755,535 | 778,957 |
| GM/GS-15 | 35 | 35 | 35 |
| GM/GS-14 | 44 | 44 | 44 |
| GM/GS-13 | 111 | 111 | 111 |
| GS-12 | 130 | 130 | 130 |
| GS-11 | 27 | 27 | 27 |
| GS-10 | 3 | 3 | 3 |
| GS-9 | 24 | 27 | 30 |
| GS-8 | 69 | 69 | 69 |
| GS-7 | 29 | 29 | 29 |
| GS-6 | 9 | 9 | 9 |
| GS-5 | 6 | 6 | 6 |
| GS-4 | 22 | 22 | 22 |
| GS-3 | 5 | 5 | 5 |
| GS-2 | 4 | 4 | 4 |
| Subtotal | 518 | 521 | 524 |
| Grades established by Act of July 1, 1944 (42 U.S.C. 207): | |||
| Director Grade | 1 | 1 | 1 |
| Senior Grade | 1 | 1 | 1 |
| Subtotal | 2 | 2 | 2 |
| Ungraded | 139 | 149 | 149 |
| Total permanent positions | 499 | 515 | 518 |
| Total positions, end of year | 664 | 677 | 680 |
| Total full-time equivalent (FTE) employment, end of year | 661 | 653 | 656 |
| Average ES salary | $146,082 | $151,107 | $155,792 |
| Average GM/GS grade | 10.9 | 10.9 | 10.9 |
| Average GM/GS salary | $73,263 | $75,783 | $78,133 |
Includes FTEs which are reimbursed from the NIH Roadmap for Medical Research
New Positions Requested
| FY 2007 Grade |
FY 2007 Number |
FY 2007 Annual Salary |
|
|---|---|---|---|
| Librarian | GS-9 | 2 | $46,247 |
| Computer Scientist | GS-13 | 1 | $79,752 |
| Total Requested | 3 |
Includes FTEs which are reimbursed from the NIH Roadmap for Medical Research
Last Reviewed: December 30, 2024