Congressional Justification FY 2021
Department of Health and Human Services
National Institutes of Health
National Library of Medicine (NLM)
2021 Congressional Justification (PDF)
FY 2021 Budget
- Organization Chart
- Appropriation Language
- Amounts Available for Obligation
- Budget Mechanism Table
- Major Changes in Budget Request
- Summary of Changes
- Budget Graphs
- Budget Authority by Activity
- Authorizing Legislation
- Appropriations History
- Justification of Budget Request
- Budget Authority by Object Class
- Salaries and Expenses
- Detail of Full-Time Equivalent Employment (FTE)
- Detail of Positions
Organizational Chart
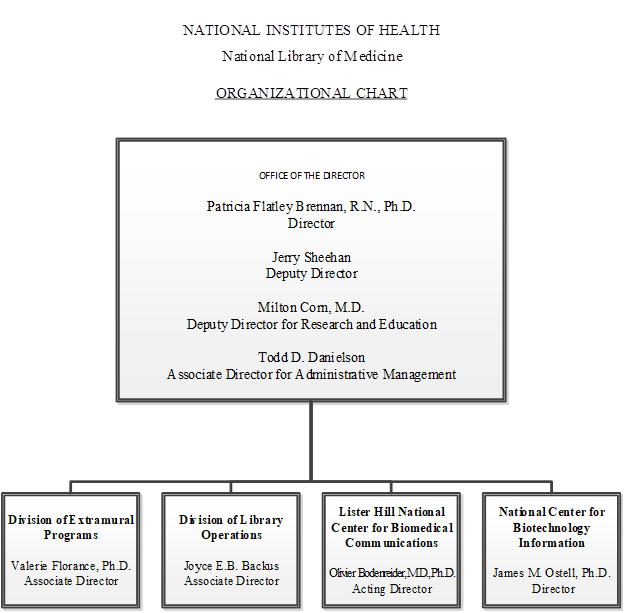
- Office of the Director
Patricia Flatley Brennan, R.N., Ph.D., Director
Jerry Sheehan, Deputy Director
Milton Corn, M.D., Deputy Director for Research and Evaluation
Todd D. Danielson, Associate Director for Administrative Management- Division of Extramural Programs
Valerie Florance, Ph.D., Director - Division of Library Operations
Joyce E. B. Backus, Associate Director - Lister Hill National Center for Biomedical Communications
Olivier Bodenreider, M.D., PhD., Acting Director - National Center for Biotechnology Information
James M. Ostell, Ph.D., Director
- Division of Extramural Programs
Appropriation Language
For carrying out section 301 and title IV of the PHS Act with respect to health information communications, [$456,911,000]$415,665,000: Provided, That of the amounts available for improvement of information systems, $4,000,000 shall be available until September 30, [2021]2022: Provided further, That in fiscal year [2020]2021, the National Library of Medicine may enter into personal services contracts for the provision of services in facilities owned, operated, or constructed under the jurisdiction of the National Institutes of Health (referred to in this title as "NIH").
Amounts Available for Obligation 1
(Dollars in Thousands)
| Source of Funding | FY 2019 Final | FY 2020 Enacted | FY 2021 President's Budget |
|---|---|---|---|
| Appropriation | $441,997 | $456,911 | $415,665 |
| Mandatory Appropriation: (non-add) | |||
| Type 1 Diabetes | (0) | (0) | (0) |
| Other Mandatory financing | (0) | (0) | (0) |
| Rescission | 0 | 0 | 0 |
| Sequestration | 0 | 0 | 0 |
| Secretary's Transfer | -1,518 | 0 | 0 |
| Subtotal, adjusted appropriation | $440,479 | $456,911 | $415,665 |
| OAR HIV/AIDS Transfers | 368 | 0 | 0 |
| HEAL Transfer from NINDS | 0 | 0 | 0 |
| Subtotal, adjusted budget authority | $440,847 | $456,911 | $415,665 |
| Unobligated balance, start of year | 2,500 | 2,500 | 0 |
| Unobligated balance, end of year | -2,500 | 0 | 0 |
| Subtotal, adjusted budget authority | $440,847 | $459,411 | $415,665 |
| Unobligated balance lapsing | -212 | 0 | 0 |
| Total obligations | $440,635 | $459,411 | $415,665 |
1 Excludes the following amounts (in thousands) for reimbursable activities carried out by this account:
FY 2019 - $7,899 FY 2020 - $7,954 FY 2021 - $7,954
Budget Mechanism Total 1
(Dollars in Thousands)
| MECHANISM | FY 2019 Final | FY 2020 Enacted | FY 2021 President's Budget | FY 2021 +/- FY 2020 Enacted | ||||
|---|---|---|---|---|---|---|---|---|
| No. | Amount | No. | Amount | No. | Amount | No. | Amount | |
| Research Projects: | ||||||||
| Noncompeting | 62 | $22,539 | 79 | $28,459 | 87 | $25,041 | 8 | -$3,418 |
| Administrative Supplements | (1) | 2 | (0) | 0 | (0) | 0 | (0) | 0 |
| Competing: | ||||||||
| Renewal | 5 | 2,658 | 2 | 1,031 | 1 | 510 | -1 | -521 |
| New | 33 | 11,473 | 28 | 8,995 | 34 | 9,691 | 6 | 696 |
| Supplements | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Subtotal, Competing | 38 | $14,131 | 30 | $10,027 | 35 | $10,201 | 5 | $175 |
| Subtotal, RPGs | 100 | $36,671 | 109 | $38,486 | 122 | $35,243 | 13 | -$3,243 |
| SBIR/STTR | 5 | 1,475 | 3 | 1,564 | 4 | 1,392 | 1 | -173 |
| Research Project Grants | 105 | $38,146 | 112 | $40,050 | 126 | $36,634 | 14 | -$3,416 |
| Research Centers: | ||||||||
| Specialized/Comprehensive | 0 | $34 | 0 | $36 | 0 | $32 | 0 | -$4 |
| Clinical Research | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Biotechnology | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Comparative Medicine | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Research Centers in Minority Institutions | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Research Centers | 0 | $34 | 0 | $36 | 0 | $32 | 0 | -$4 |
| Other Research: | ||||||||
| Research Careers | 12 | $1,799 | 13 | $1,719 | 5 | $596 | -8 | -$1,123 |
| Cancer Education | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Cooperative Clinical Research | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Biomedical Research Support | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Minority Biomedical Research Support | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Other | 44 | 24,773 | 41 | 24,704 | 42 | 23,144 | 1 | -1,561 |
| Other Research | 56 | $26,572 | 54 | $26,423 | 47 | $23,739 | -7 | -$2,684 |
| Total Research Grants | 161 | $64,752 | 166 | $66,510 | 173 | $60,406 | 7 | -$6,104 |
| Ruth L Kirchstein Training Awards: | FTTPs | FTTPs | FTTPs | FTTPs | ||||
| Individual Awards | 8 | $342 | 9 | $386 | 7 | $301 | -2 | -$85 |
| Institutional Awards | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Total Research Training | 8 | $342 | 9 | $386 | 7 | $301 | -2 | -$85 |
| Research & Develop. Contracts | 0 | $18 | 0 | $18 | 0 | $17 | 0 | -$2 |
| (SBIR/STTR) (non-add) | (0) | (15) | (0) | (18) | (0) | (17) | (0) | (-2) |
| Intramural Programs | 575 | 355,960 | 638 | 369,381 | 638 | 335,356 | 0 | -34,024 |
| Res. Management & Support | 84 | 19,775 | 103 | 20,616 | 103 | 19,586 | 0 | -1,031 |
| Res. Management & Support (SBIR Admin) (non-add) | (0) | (2) | (0) | (0) | (0) | (0) | (0) | (0) |
| Construction | 0 | 0 | 0 | 0 | ||||
| Buildings and Facilities | 0 | 0 | 0 | 0 | ||||
| Total, NLM | 659 | $440,847 | 741 | $456,911 | 741 | $415,665 | 0 | -$41,246 |
1 All items in italics and brackets are non-add entries.
Major Changes in the Fiscal Year 2021 President’s Budget Request
Major changes in the FY 2021 President’s Budget request for the National Library of Medicine (NLM), by budget mechanism and/or activity detail are briefly described below. Note that there may be overlap between budget mechanism and activity detail; thus, these highlights will not sum to the total change for NLM’s FY 2021 President’s Budget request, which is $415.7 million, a decrease of $41.2 million from the FY 2020 Enacted level. The FY 2021 President's Budget reflects the Administration's fiscal policy goals for the Federal Government. Within that framework and informed by the NLM Strategic Plan, 2017-2027, the NIH Strategic Plan for Data Science, and the 2018 Blue Ribbon Panel review of NLM’s Intramural Research Program, NLM will pursue its highest research priorities through strategic investments and careful stewardship of appropriated funds.
Extramural Programs (-$6.2 million; total $60.7 million): With this level of funding, NLM will support the same number of university-based biomedical informatics and data science training programs as under the FY 2020 Enacted level with a reduced funding level. NLM will award an estimated 35 new research project grants in biomedical informatics and data science and will support the outreach and stakeholder engagement efforts of the National Network of Libraries of Medicine at a reduced level of funding.
Intramural Programs (-$34.0 million; total $335.4 million): NLM’s intramural programs encompass both intramural research and information service programs that support advances in computational health information sciences; development of advanced biomedical information systems, standards, and research tools; acquisition, storage and distribution of biomedical data; and delivery of reliable, high-quality information services. NLM will maintain funding for intramural research and prioritize activities that support NIH-wide interests in data science. It will continue to seek ways to streamline and improve the efficiency of its information services while maintaining support for mission critical systems that are heavily used by researchers, clinicians, and the general public. NLM will continue its role as a central coordinating body for the Department of Health and Human Services as relates to standard clinical vocabularies while seeking additional efficiencies in these programs.
Summary of Changes
(Dollars in Thousands)
| Budget Category | Amount |
|---|---|
| FY 2020 Enacted | $456,911 |
| FY 2021 President's Budget | $415,665 |
| Net change | -$41,246 |
| CHANGES | FY 2021 President's Budget | Change from FY 2020 Enacted | ||
|---|---|---|---|---|
| FTEs | Budget Authority | FTEs | Budget Authority | |
| A. Built-in: | ||||
| 1. Intramural Programs: | ||||
| a. Annualization of January 2020 pay increase & benefits | $114,689 | $731 | ||
| b. January FY 2021 pay increase & benefits | 114,689 | 1,761 | ||
| c. Paid days adjustment | 114,689 | -428 | ||
| d. Differences attributable to change in FTE | 114,689 | 0 | ||
| e. Payment for centrally furnished services | 3,896 | -205 | ||
| f. Cost of laboratory supplies, materials, other expenses, and non-recurring costs | 216,772 | 3,468 | ||
| Subtotal | $5,327 | |||
| 2. Research Management and Support: | ||||
| a. Annualization of January 2020 pay increase & benefits | $12,521 | $80 | ||
| b. January FY 2021 pay increase & benefits | 12,521 | 190 | ||
| c. Paid days adjustment | 12,521 | -47 | ||
| d. Differences attributable to change in FTE | 12,521 | 0 | ||
| e. Payment for centrally furnished services | 787 | -41 | ||
| f. Cost of laboratory supplies, materials, other expenses, and non-recurring costs | 6,278 | 83 | ||
| Subtotal | $265 | |||
| Subtotal, Built-in | $5,592 | |||
| FY 2021 President's Budget | Change from FY 2020 Enacted | |||
|---|---|---|---|---|
| CHANGES | No. | Amount | No. | Amount |
| B. Program: | ||||
| 1. Research Project Grants: | ||||
| a. Noncompeting | 87 | $25,041 | 8 | -$3,418 |
| b. Competing | 35 | 10,201 | 5 | 175 |
| c. SBIR/STTR | 4 | 1,392 | 1 | -173 |
| Subtotal, RPGs | 126 | $36,634 | 14 | -$3,416 |
| 2. Research Centers | 0 | $32 | 0 | -$4 |
| 3. Other Research | 47 | 23,739 | -7 | -2,684 |
| 4. Research Training | 7 | 301 | -2 | -85 |
| 5. Research and development contracts | 0 | 17 | 0 | -2 |
| Subtotal, Extramural | $60,723 | -$6,191 | ||
| FTEs | FTEs | |||
| 6. Intramural Programs | 638 | $335,356 | 0 | -$39,351 |
| 7. Research Management and Support | 103 | 19,586 | 0 | -1,296 |
| 8. Construction | 0 | 0 | ||
| 9. Buildings and Facilities | 0 | 0 | ||
| Subtotal, Program | 741 | $415,665 | 0 | -$46,838 |
| Total changes | -$41,246 | |||
Fiscal Year 2021 Budget Graphs
History of Budget Authority and FTEs:
Bar Graph for Funding Levels by Fiscal Year for FY2017 through FY2021
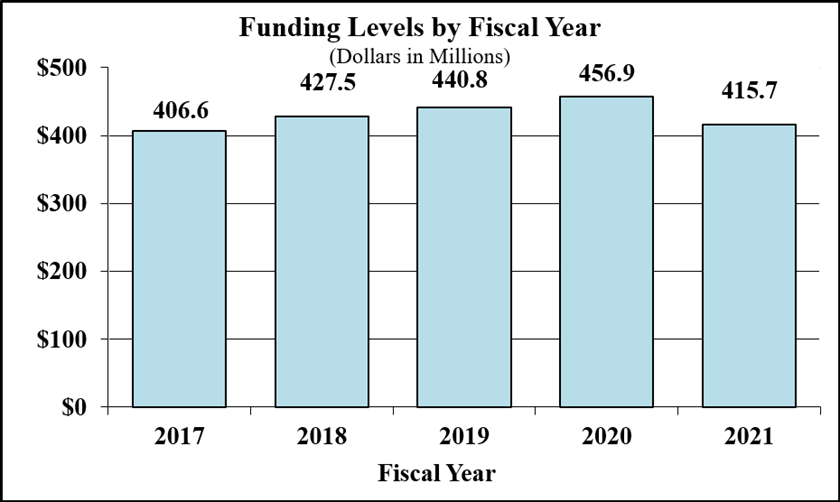
Data for Funding Levels by Fiscal Year for FY2017 through FY2021
| Fiscal Year | Funding (Dollars in Millions) |
|---|---|
| 2017 | 406.6 |
| 2018 | 427.5 |
| 2019 | 440.8 |
| 2020 | 456.9 |
| 2021 | 415.7 |
Bar Graph of FTEs by Fiscal Year for FY2017 through FY2021
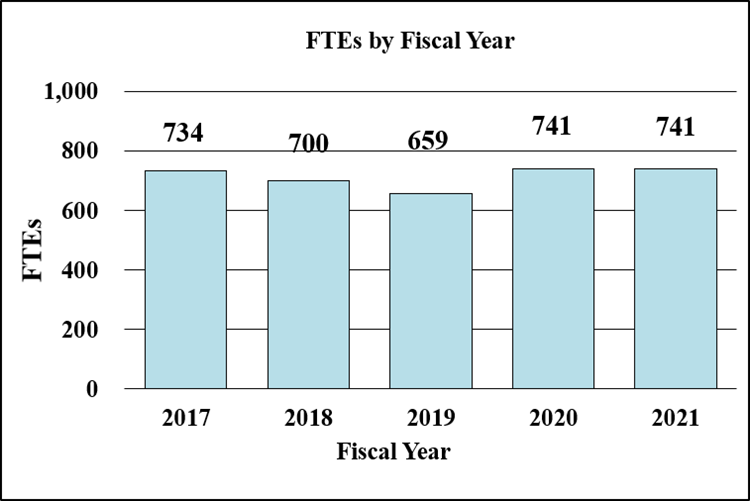
Data for FTEs by Fiscal Year for FY2017 through FY2021
| Fiscal Year | FTEs |
|---|---|
| 2017 | 734 |
| 2018 | 700 |
| 2019 | 659 |
| 2020 | 741 |
| 2021 | 741 |
Distribution by Mechanism:
Pie Chart of FY2021 Budget Mechanisms
(Budget in Thousands)
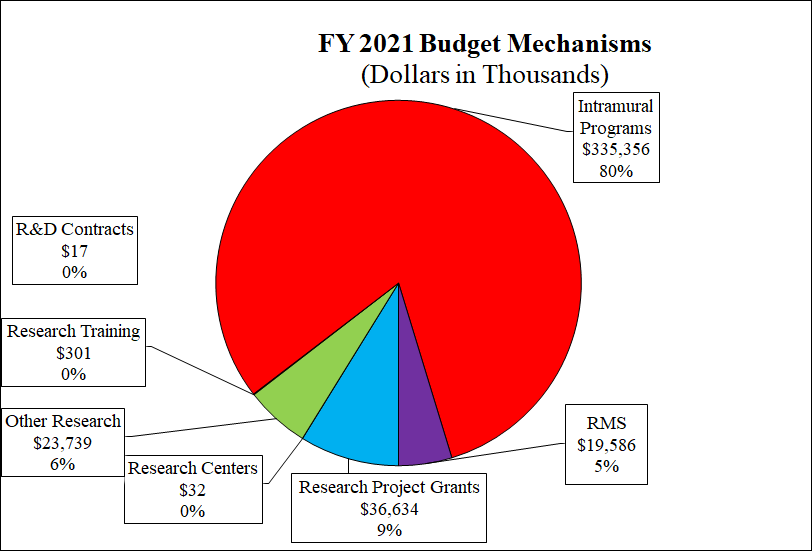
Data for FY2021 Budget Mechanisms
(Budget in Thousands)
| Mechanism | Dollars | Percentage |
|---|---|---|
| Research Project Grants | $36,634 | 9% |
| Research Centers | $32 | 0% |
| Other Research | $23,739 | 6% |
| Research Training | $301 | 0% |
| R&D Contracts | $17 | 0% |
| Intramural Programs | $335,356 | 80% |
| RMS | $19,586 | 5% |
Bar Graph of FY2021 Estimated Percent Change from FY 2020 Mechanism
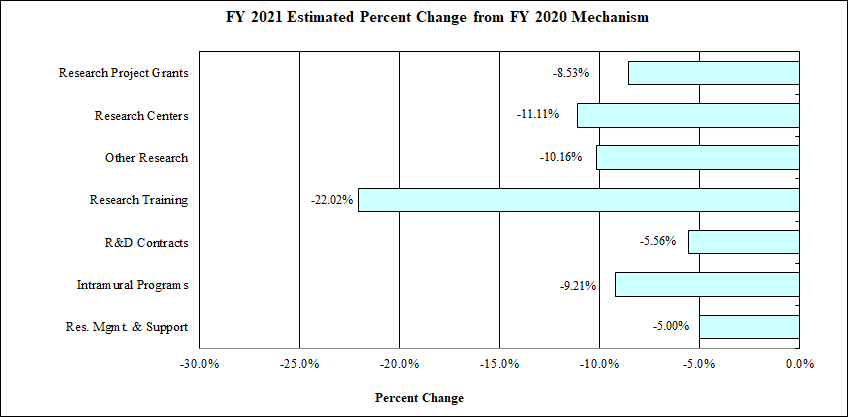
Data for Bar Graph of FY2021 Estimated Percent Change from FY 2020 Mechanism
| Mechanism | Percentage Change |
|---|---|
| Research Project Grants | -8.5% |
| Research Centers | -11.1% |
| Other Research | -10.2% |
| Research Training | -22.0% |
| R&D Contracts | -5.6% |
| Intramural Programs | -9.2% |
| Res. Mgmt. & Support | -5.0% |
Budget Authority By Activity 1
(Dollars in Thousands)
| FY 2019 Final | FY 2020 Enacted | FY 2021 President's Budget | FY 2021 +/- FY2020 | |||||
|---|---|---|---|---|---|---|---|---|
| Extramural Research | FTE | Amount | FTE | Amount | FTE | Amount | FTE | Amount |
| Detail | ||||||||
| Health Information for Health Professionals and the Public (NN/LM) | $11,270 | $11,132 | $12,000 | $868 | ||||
| Informatics Resources for Biomedicine and Health | 15,696 | 15,732 | 12,089 | -3,643 | ||||
| Biomedical Informatics Research | 38,146 | 40,050 | 36,634 | -3,416 | ||||
| Subtotal, Extramural | $65,113 | $66,914 | $60,723 | -$6,191 | ||||
| Intramural Programs | 575 | $355,960 | 638 | $369,381 | 638 | $335,356 | 0 | -$34,024 |
| Research Management & Support | 84 | $19,775 | 103 | $20,616 | 103 | $19,586 | 0 | -$1,031 |
| TOTAL | 659 | $440,847 | 741 | $456,911 | 741 | $415,665 | 0 | -$41,246 |
1Includes FTEs whose payroll obligations are supported by the NIH Common Fund.
Authorizing Legislation
| PHS Act/ Other Citation |
U.S. Code Citation |
2020 Amount Authorized |
FY 2020 Enacted | 2021 Amount Authorized |
FY 2021 President's Budget | |
|---|---|---|---|---|---|---|
| Research and Investigation | Section 301 | 42§241 | Indefinite | $456,911,000 | Indefinite | $415,665,000 |
| National Library of Medicine | Section 401(a) | 42§281 | Indefinite | Indefinite | ||
| Total, Budget Authority | $456,911,000 | $415,665,000 | ||||
Appropriations History
| Fiscal Year | Budget Estimate to Congress | House Allowance | Senate Allowance | Appropriation |
|---|---|---|---|---|
| 2012 | $387,153,000 | $387,153,000 | $358,979,000 | $338,278,000 |
| Rescission | $639,345 | |||
| 2013 | $372,651,000 | $381,981,000 | $337,638,655 | |
| Rescission | $675,277 | |||
| Sequestration | ($16,947,139) | |||
| 2014 | $382,252,000 | $387,912,000 | $327,723,000 | |
| Rescission | $0 | |||
| 2015 | $372,851,000 | $336,939,000 | ||
| Rescission | $0 | |||
| 2016 | $394,090,000 | $341,119,000 | $402,251,000 | $394,664,000 |
| Rescission | $0 | |||
| 2017¹ | $395,684,000 | $407,086,000 | $412,097,000 | $407,510,000 |
| Rescission | $0 | |||
| 2018 | $373,258,000 | $413,848,000 | $420,898,000 | $428,553,000 |
| Rescission | $0 | |||
| 2019 | $395,493,000 | $433,671,000 | $442,230,000 | $441,997,000 |
| Rescission | $0 | |||
| 2020 | $380,463,000 | $463,599,000 | $465,837,000 | $456,911,000 |
| Rescission | $0 | |||
| 2021 | $415,665,000 |
1 Budget Estimate to Congress includes mandatory financing.
Justification of Budget Request
Authorizing Legislation: Section 301 and title IV of the Public Health Service Act, as amended.
Budget Authority (BA):
| FY 2019 Final |
FY 2020 Enacted |
FY 2021 President's Budget |
FY 2021 +/- FY 2020 |
|
|---|---|---|---|---|
| BA | $440,847,000 | $456,911,000 | $415,665,000 | -$41,246,000 |
| FTE | 659 | 741 | 741 | 0 |
Program funds are allocated as follows: Competitive Grants/Cooperative Agreements; Contracts; Direct Federal/Intramural and Other.
Director's Overview
The National Library of Medicine (NLM) is a leader in research in biomedical informatics,1 data science, and open science. It is also the world’s largest biomedical library. NLM’s research and information services advance digitization in biomedicine to support scientific discovery and health care. NLM pioneers new ways to make biomedical data and information more accessible to those who need it, building tools for better data management and personal health, creating a more diverse, data-skilled workforce, and engaging with stakeholders. In these ways, NLM’s activities support a wide array of stakeholders, including researchers, clinicians, librarians, public health officials, other Federal agencies, and the public.
NLM's cutting-edge research and training programs, with a focus on artificial intelligence (AI), machine learning, computational biology, and health data standards, catalyze basic biomedical science, data-driven discovery, and health care delivery. Recent applications of NLM research have enhanced natural language processing methods to understand medical text and improve detection of macular degeneration and diagnosis of early-stage cervical cancer. NLM research on health data standards makes it possible to rapidly exchange and use data to improve health and advance biomedical research. NLM research has created new computational approaches to comparative genomics to predict previously unknown protein and gene functions, classify and characterize bacterial toxins, and better understand the roles of individual proteins in the maintenance and function of chromosomes. NLM also develops and deploys novel approaches to data processing to rapidly evaluate, annotate, and publish sequences of the influenza virus (commonly called the flu); makes huge datasets quickly available for disease outbreak analysis; and develops tools to make infectious disease data easier to find and understand.
1 https://www.amia.org/about-amia/science-informatics
NLM develops and applies innovative approaches to acquire, organize, curate, and deliver current biomedical information across the United States and around the globe. NLM’s advanced biomedical information services are among the most visited websites in the Federal Government, providing researchers, health care professionals, and the public with access to high quality biomedical information and data, including biomedical literature, genomic data, clinical trial data, and chemical information. NLM supports data-driven discovery for better outcomes in health care and public health by improving data collection and sharing for minority health, women’s health, health equity, and health quality, including multilingual health information libraries and innovative clinical records integration for youth emancipating from foster care. NLM’s biomedical information systems accelerate science, broaden opportunities for collaboration, provide platforms for private sector innovation, and increase the return on investments in research. NLM participates in NIH-wide efforts to foster a culture that advances science and ensures the development and retention of a diverse, safe, and inclusive workforce.
Twenty-year Retrospective and Today: Over the last 20 years, NLM has been a key driver of research advances in biomedical informatics, data science, and open science. Data science is the development and application of novel approaches, processes, and systems to extract knowledge and insights from increasingly large and/or complex sets of data. Open science accelerates biomedical research and discovery by ensuring the products and processes of scientific research are readily available, accessible, and usable by the broadest set of potential users. Long before data science was recognized as a field of study, NLM scientists devised innovative ways to manage and analyze large datasets and rapidly discover novel associations and new interpretations. NLM’s Basic Local Alignment Search Tool (BLAST), for example, is one of the most powerful and popular biological sequence analysis tools available in the public domain and has kept pace with the substantial and increasing amounts of sequence data being generated. NLM recently moved BLAST to modern computer platforms, i.e., “the cloud,” enhancing opportunities for research and discovery.
NLM was an early adopter of the Internet, providing free online public access to biomedical literature more than 20 years ago through Grateful Med, a tool for retrieving key references from MEDLINE, NLM’s database of biomedical citations and abstracts. NLM has continued to modernize its platforms and accelerate access to growing volumes of biomedical data and information via the Internet. With the launch of PubMed in 1996, NLM solidified its role as a leader in providing free, internet-based access to biomedical data and information. PubMed has become the most heavily used biomedical literature citation database in the world, providing access to MEDLINE and other citation information. NLM’s PubMed Central (PMC), established in 2000 as an electronic archive of peer-reviewed biomedical and life sciences literature, now provides researchers, clinicians, and the public with access to the full text of more than 5.5 million articles. It serves as the repository for NIH’s Public Access Policy, which ensures access to articles summarizing the results of NIH-funded research. With the explosion of genomic information over the last 20 years, NLM developed a family of online databases, for example GenBank, Reference Sequence database (RefSeq), and database of Genotypes and Phenotypes (dbGaP), for scientists to further explore genetic information without the need to re-sequence biological samples. These databases have advanced to keep pace with the increase in volume of genomic data. NLM is now taking steps to move some of these data to the cloud to enable large-scale computing (see Program Portrait 1, page 14).
Over the last 20 years, NLM has also accelerated efforts to improve transparency of, and access to, clinical trials information. NLM’s ClinicalTrials.gov, launched in 2000, is the world’s largest publicly accessible database of privately and publicly funded clinical studies, including NIH-funded clinical trials. It supports Federal laws, as well as policies implemented in the United States and internationally. By the end of FY 2019, ClinicalTrials.gov contained information on more than 318,000 registered studies, 39,000 of which include study results and adverse events information.
In addition, NLM has laid the groundwork for the principled exchange of health data for clinical and research purposes. As the coordinating body for clinical terminology standards within the Department of Health and Human Services (HHS), NLM led Federal efforts to drive broad development and adoption of standards that facilitate exchanging and combining health data. NLM served as a leader across NIH and the Federal Government to address challenges to make the rapidly growing amounts of biomedical research data more accessible and usable. Through these efforts, NLM played a key role in developing consistent policies and encouraging best practices in managing and sharing federally-funded research data. It continues to participate in efforts which investigate the re-use of biomedical datasets, develop related metrics, identify incentives for data scientists to focus on biomedical challenges, and ensure sustainability of an open, digital ecosystem for biomedical research.
From FY 2015-2019 NLM expanded investments in research, training, and community engagement, as well as the physical and technical infrastructure for its information services. During this period, NLM nearly doubled the number of research project grants awarded, supported more early-stage investigators (i.e., scientists early in their career), and provided NLM scientists with new opportunities in data science, informatics, health data standards, and computational biology. It expanded biomedical data science training and resources for undergraduate, graduate, and postgraduate students. NLM also expanded the amount of high-quality information, tools, and data available to researchers, health professionals, policy makers, and the public. NLM addressed critical needs for improving the stability and security of its information systems and associated technical infrastructure, efficiencies in cataloging and performance measurement, and physical infrastructure to ensure a safe and productive work environment.
Investing for the Future: NLM will continue making biomedical information and knowledge accessible to more people in more ways by expanding research done by NLM’s intramural and extramural scientists in data science and biomedical informatics. NLM will facilitate the availability and use of data through methods research, support for open science, a data-skilled biomedical workforce, a modern information technology (IT) infrastructure, enhanced dissemination and engagement, and responsible use of data for AI. It is systematically evaluating and strengthening the technical infrastructure for its public-facing websites and applications by unifying platforms and balancing local storage with cloud-based solutions.
Program Portrait 1: Genomic Sequence Data Available in the Cloud
NLM’s Sequence Read Archive (SRA) is the world’s largest publicly available repository of next-generation genome sequence data, with more than 9 million records comprising 25 petabytes of data. More than 100,000 users each month download subsets of SRA data for analysis in their local computing environments. To improve access and utility of SRA data, NLM launched a major effort in FY 2019, to upload the public access SRA data to two commercial clouds that have agreements with NIH’s Science and Technology Research Infrastructure for Discovery, Experimentation, and Sustainability (STRIDES) Initiative. This transition significantly expands the discovery potential of the data. Freed from the limitations of local storage and computational resources, users are empowered to compute across the full corpus of SRA data without having to download and store large volumes of data. Moving to cloud platforms also makes it possible to develop customized tools and methods for asking research questions of the data. Publicly accessible SRA data include genomes of viruses, bacteria, and nonhuman higher organisms, as well as gene expression data, metagenomes, and a small amount of human genomic data that is consented to be publicly available (e.g., data from the 1000 Genomes Project). NLM is working to make SRA’s controlled-access human genomic data available on commercial cloud platforms, with higher levels of security and oversight, to ensure continued protection of data from human samples and appropriate authorization and authentication of users of these data. NLM will facilitate NIH’s strategic efforts to move these valuable datasets into commercial cloud computing platforms through technical guidance to enact privacy-preserving policies for controlling access, including a strategic vision for identity management anchored on centralized authentication and authorization for access to datasets from across NIH Institutes and Centers (ICs).
In FY 2019, NLM supported several training and outreach activities to extend NLM’s reach and engage talent, including:
- Research training programs in biomedical and public health informatics and data science at 16 universities that supported 200 predoctoral and postdoctoral fellows. To help attract members of underrepresented populations, NLM provided supplements for collaboration between NLM’s university-based training programs and minority-serving institutions to share curricula and course materials in biomedical data science.
- Joint initiatives with the National Science Foundation (NSF) to engage more scientists with expertise in data science research with biomedical researchers. In FY 2019, NLM awarded 10 grants, 5 each in two different NLM-NSF initiatives, bringing 9 new early-stage investigators (10 years or less since their terminal degree) to NIH.
- Training courses for nearly 50 NIH intramural scientists, as well as opportunities for fellows co-funded by other NIH ICs, to apply informatics science to research, education, and clinical care. NLM aims to expand intramural training in biomedical informatics and data science and improve outreach to recruit diverse and highly-qualified trainees.
- Participation in NIH’s Pathways Program to recruit interns and recent graduates from diverse populations.
- Implementation of the NIH Strategic Plan for Data Science tactic focused on data science training for NIH staff. NLM spearheaded a needs assessment of extramural program officers to inform training initiatives on data science and open science and provided training in data science and open science to NLM staff.
- Preparation of information professionals for data-driven discovery through NLM’s National Network of Libraries of Medicine (NNLM). In FY 2019, more than 32,000 attendees participated in NNLM data-related classes and events.
- Management of citizen science efforts to improve and apply NLM products and services. NLM led 19 code-a-thons (i.e., hands-on, team-based training projects) with approximately 900 external participants, investigating topics ranging from antimicrobial resistance to discovery of novel splice isoforms in RNA-sequence data using NLM’s RefSeq collection.
Overall Budget Policy
The FY 2021 President’s Budget request is $415.7 million, a decrease of $41.2 million from the FY 2020 Enacted level for NLM. Funds are included to sustain intramural research in computational health information sciences and methods, such as AI and machine learning, and maintain NLM’s most heavily used information services, including those that provide access to published biomedical literature and consumer health information. At this level, support for NLM databases and services will be prioritized, ensuring mission critical support for NIH data science goals and data sharing policies. NLM will retain key intramural data science activities, including processing and organizing genomic data resulting from NIH-wide investments in whole genome sequencing; evaluation of microbial and viral pathogens to support outbreak detection and response by the Centers for Disease Control and Prevention (CDC), Food and Drug Administration (FDA), and Department of Agriculture (USDA); enhancing registration and results reporting in ClinicalTrials.gov; and updating and disseminating clinical terminology standards required for interoperability of U.S. health data. NLM will also award up to 35 new research project grants through its extramural programs. NLM will continue to leverage the National Network of Libraries of Medicine to advance community engagement in the NIH All of Us initiative and will prioritize and consolidate outreach programs that promote access and training in the effective use of NLM resources for scientists, clinicians, patients, and the general public.
Program Descriptions and Accomplishments
Intramural Programs
NLM’s intramural programs encompass two major activities: 1) Intramural Research and Training; and 2) Biomedical Information Services.
Intramural Research and Training: NLM conducts research and training in computational health science, which includes biomedical informatics, computational biology, AI, machine learning, and deep learning. This work generates new knowledge, methods, and tools that support biomedical discovery, image interpretation, and clinical records analysis to enable scientists and clinicians to better understand basic biology, discover new approaches to care, and improve clinical decisions and health.
Recent NLM research has resulted in scientific advances that:
- Helped quantify horizontal gene transfer, a recently discovered biological mechanism that explains species diversity, and explored how stalling in the metabolic process disrupts the function of transfer RNA and alters subsequent protein function. NLM also developed new computational approaches to classify and characterize bacterial toxins.
- Contributed to the comprehensive characterization of proteins involved in the functions of clustered regularly spaced short palindromic repeats (CRISPR) systems, which enable genome editing. This work was part of a larger effort to predict the functions of proteins of biological importance. The majority of the predicted biochemical activities and biological functions have been successfully validated experimentally through collaborations with several laboratories, as well as in independent studies.
- Discovered new families of viruses and predicted the functions of their genes. The research included description of a large, previously unknown family of bacteriophages (viruses that infect bacteria) that includes cross-assembly phage (crAssphage), the most abundant virus in the human microbiome. These findings are crucial for the further characterization of the human microbiome and have already stimulated numerous follow-up studies.
- Laid the groundwork for a new tool, called single-cell sub-Populations Comparison, that helps researchers understand the differences between populations of cells from single-cell experiments, a research area of heightened interest. The tool allows researchers to define both the common and unique cell types across many experiments. Scientists can use it to identify similar and distinct cell types present in single-cell experiments, including cells with different disease status, different developmental stages, and different sexes and species.
NLM research has also resulted in novel computational approaches for analyzing whole genome sequencing data from bacteria to help quickly resolve outbreaks of foodborne illness by identifying the bacterial pathogens involved and helping to trace them back to their source. These approaches are embedded in NLM’s Pathogen Detection Pipeline, which has supported more than 370 outbreak investigations to date. In addition, NLM research has developed new approaches to data processing to rapidly evaluate, annotate, and disseminate high-quality influenza sequence data, making large, well-organized datasets quickly available to scientists seeking to define factors that impact influenza virus outbreak. More than 790,000 influenza genome sequences are available, with almost 110,000 submitted in FY 2019. NLM is also contributing to research on genetic causes of antimicrobial resistance (AMR) as part of an ongoing collaboration with the National Antimicrobial Resistance Monitoring System, a public health surveillance system that tracks changes in the antimicrobial susceptibility of foodborne pathogens. Using phenotypes obtained by other Federal agencies (i.e., CDC, FDA, and USDA) for more than 6,000 pathogen isolates, NLM researchers validated the AMRFinder tool they built for antimicrobial resistance gene and protein predictions. This work resulted in greater than 98 percent consistent predictions. The tool also predicts the genes and proteins in the n ew National Database of Antibiotic Resistant Organisms which consists of more than 400,000 pathogens and is part of the National Action Plan for Combating Antibiotic-resistant Bacteria.
NLM research also develops and applies sophisticated analytical approaches to the study of clinical phenomena, such as the effects of medications on health outcomes, using large clinical datasets. In FY 2019, NLM researchers used such techniques to develop insights into the relationship and risk of seven antibiotics, called fluoroquinolones, for tendon rupture in a cohort of 1.2 million Medicare enrollees, and the effects of estrogens on patient survival in a large population of postmenopausal women. NLM researchers used structured clinical terminologies and natural language processing to recognize complex clinical phenomena and guide clinical decisions for precision medicine in oncology. Using simulated clinical cases created by precision oncologists to describe a patient’s cancer type, relevant genetic variants, and demographic information, NLM researchers evaluated tools to support precision medicine by retrieving journal articles about treatments relevant to specific patients and finding clinical trials (from ClinicalTrials.gov) for potential enrollment. NLM enhances the discoverability of biomedical literature by developing and disseminating open source software tools to support timely indexing of the literature with key terms from structured terminologies.
NLM research results in novel AI algorithms for analyzing medical images that can improve detection, diagnosis, and treatment of disease. Much of this work involves collaboration with other NIH ICs with clinical expertise that complements NLM’s informatics and data science expertise, resulting in advances such as:
- A screening tool for cervical cancer, developed in partnership with the National Cancer Institute (NCI) and private industry, for improving classification of cervical cancer and assisting in early treatment in low-resource areas. The tool, which is being developed for mobile phones, augments human expertise and understands disease expression (e.g. variability in age, ethnicity, and disease severity).
- Novel approaches, developed in collaboration with the National Eye Institute, to classify the severity of age-related macular degeneration (AMD) and predict risk of progression to late-stage AMD better than existing clinical standards.
- An algorithm improved and applied in collaboration with NCI and global private-public partners, to detect abnormalities in chest X-ray images and screen for tuberculosis in low-resource settings for populations with high incidence of HIV.
- An algorithm, developed in collaboration with the National Institute of Allergy and Infectious Diseases, that screens for malaria with 99 percent accuracy by detecting the presence of the malaria-causing parasite in red blood cell images. Recent work uses three- dimensional images of thick blood smears to capture the entire volume of a single drop of blood, increasing the likelihood of finding parasites in patients with low parasite counts. The smartphone-based algorithm underwent successful field tests in low-resource areas and has broad applicability for recognizing abnormal blood cells.
NLM’s intramural research programs provide research training in biomedical informatics, clinical informatics, and computational biology for post-doctoral trainees, along with high school, college, and graduate students, medical professionals, and visiting scientists. Trainees participate in NLM research projects on medical imaging, AI, machine learning, medical terminology, and other cutting-edge research. They apply computational tools to research problems in molecular and structural biology, genetics, genomics, proteomics, phylogeny, and related fields. In FY 2019, NLM supported nearly 50 intramural research fellows in short-term and multi-year on-campus training.
Program Portrait 2: Fostering the Biomedical Informatics and Data Science Workforce
NLM is committed to building a diverse workforce to meet the challenges of data-powered discovery by ensuring proficiency in data science and open science, expanding research methods that support rigor and reproducibility, and engaging the next generation of researchers. NLM funds Ph.D.-level research training in biomedical informatics and data science and partners across NIH to ensure inclusion of data science and open science core skills in all NIH training programs. Through training and resource grants, NLM supports the training of librarians, information science professionals, and other research facilitators, so they can manage a data-driven future. To increase diversity in the fields it supports, NLM partners with high schools, information schools, and minority-serving institutions.
Biomedical Information Services: Many of NLM’s intramural programs support the development and operation of heavily used biomedical information services, such as PubMed, ClinicalTrials.gov, and dbGaP. Each day, more than 6 million people use NLM websites and download 115 terabytes of data. Thousands of researchers and businesses submit 15 terabytes of data daily. Annually, NLM information systems process more than six billion human interactions and eight billion computer-to-computer interactions. NLM continually expands biomedical information services to accommodate a growing volume of relevant data and information and enhances these services to support research and discovery.
In FY 2019, NLM made numerous enhancements to its information services for biomedical literature, clinical trials and consumer health information, and molecular biology and bioinformatics:
- Biomedical Literature: NLM added 1.35 million citations to PubMed, its database of citations and abstracts to the biomedical literature, increasing the total content to more than 30 million citations. NLM also launched a new PubMed platform for an improved user experience, including a new search algorithm with relevance rankings and better tools for citations. In , NLM added 600,000 full-text articles to PMC and continued linking articles to associated data by aggregating data citations, data availability statements, and supplementary materials. Since featuring these links more prominently, daily downloads of supplementary material have increased by 30 percent. Of the more than 5.5 million articles in PMC, a subset of about 3 million articles is available for bulk retrieval for text mining and other research purposes. Ten other Federal agencies use PMC as the repository for publications collected under their public access policies to ensure free public access to the results of taxpayer-funded research.
- Clinical Trials and Consumer Health Information: In FY 2019, NLM added 32,000 new clinical research studies and 6,100 new results summaries to ClinicalTrials.gov, the world’s largest publicly accessible database of privately and publicly funded clinical studies. NLM initiated a long-term project to modernize ClinicalTrials.gov to deliver on a flexible, extensible, scalable, and sustainable platform that will accommodate growth and improve efficiency consistent with applicable laws and policies. MedlinePlus had more than 300 million users who accessed its trusted, authoritative consumer health information in FY 2019. MedlinePlus offers information for patients and families on a broad variety of health conditions, along with thousands of links to other reputable sources of information. Content is provided in English and Spanish. NLM’s MedlinePlus Connect service works with electronic health records (EHRs), patient portals, and other healthcare IT systems to deliver information from MedlinePlus to patients and providers at the point of need. In FY 2019, MedlinePlus Connect responded to 252 million requests from healthcare IT systems.
- Molecular Biology and Bioinformatics Resources: In addition to SRA and BLAST, this vast array of resources includes more than 40 integrated molecular biology databases and bioinformatics software tools such as GenBank, RefSeq, dbGaP, and ClinVar. In FY 2019, NLM added 380 million sequences to GenBank, the database of all publicly available genetic sequences, and improved the quality and accessibility of viral sequence data. It added nearly 42 million sequence records to RefSeq (a 26 percent increase), which provides a comprehensive collection of sequence and gene information as well as an integrated and well-annotated view of the genetic elements contributing to the nature and behavior of all studied organisms (e.g., human, model organisms, microbes, and viruses). It also facilitated the submission of 200 studies to dbGaP, which supports NIH’s Genomic Data Sharing Policy and provides archival and access services for large-scale human genomic data, enabling the data to be analyzed by other investigators. By the end of FY 2019, dbGaP contained data from 1,200 studies, and more than 2,000 research papers had been published based on new analyses of these data. NLM also processed more than 179,000 submissions in its ClinVar database, bringing the total to over 880,000 submissions. ClinVar aggregates information about human genomic variations, their clinical significance, and their relationship to human health, providing important information to the genetic testing and clinical communities.
- NLM’s Historical Digital Collections: This free online repository of historical and modern biomedical resources includes books, manuscripts, still images, videos, sound recordings, and maps serving more than 180,000 librarians, researchers, and others annually.
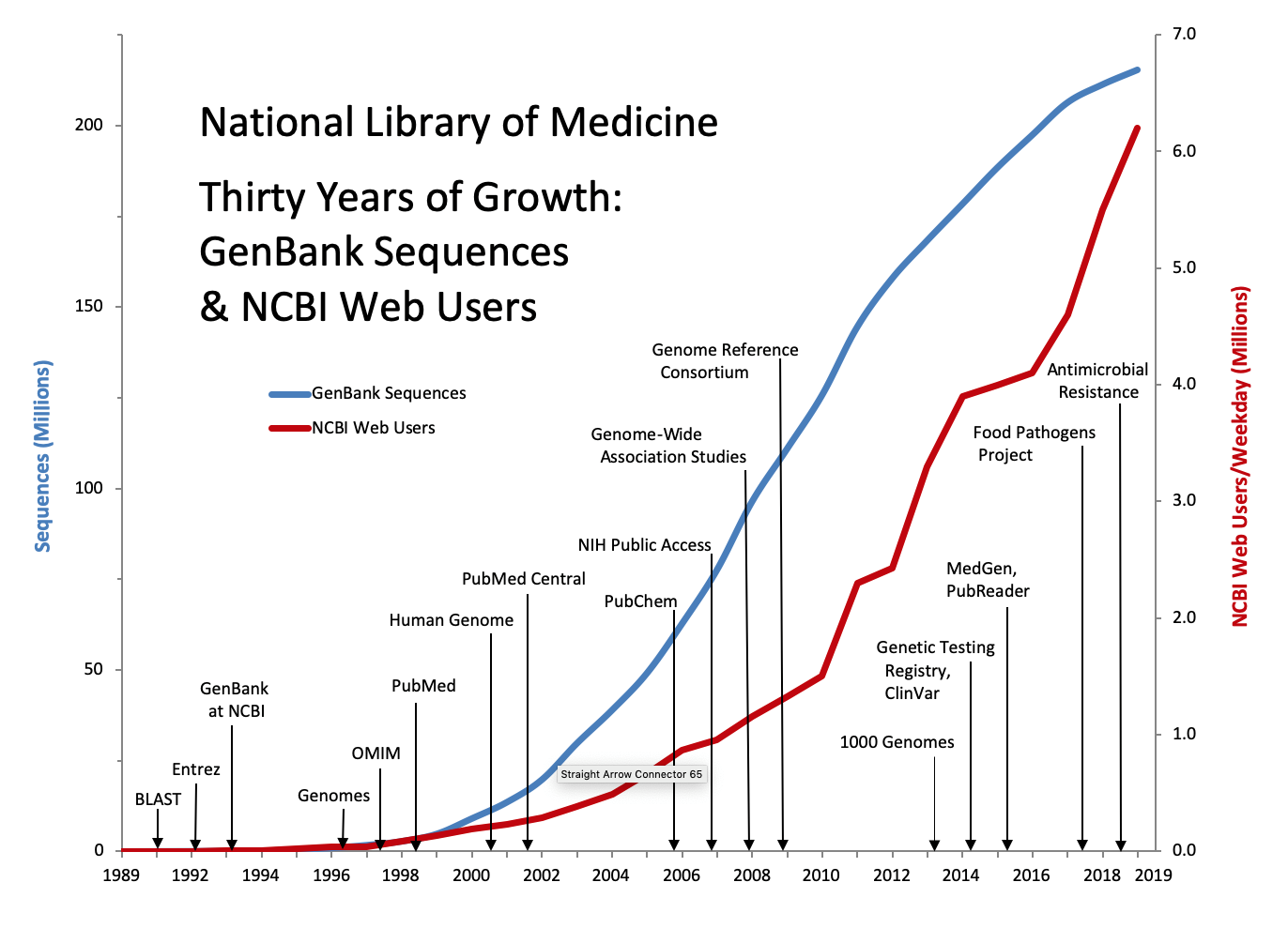
Data for Twenty Nine Years of Growth: NCBI Data and User Services
| Year | Product Release | GenBank Sequences | Users (Average) |
|---|---|---|---|
| 1989 | 30,010 | 1,200 | |
| 1990 | BLAST | 40,295 | 1,500 |
| 1991 | Entrez | 58,952 | 2,700 |
| 1992 | GenBank at NCBI | 87,846 | 3,200 |
| 1993 | 143,492 | 4,500 | |
| 1994 | 215,273 | 9,000 | |
| 1995 | Genomes | 555,694 | 19,000 |
| 1996 | OMIM | 1,021,211 | 36,000 |
| 1997 | PubMed | 1,765,847 | 39,268 |
| 1998 | 2,837,897 | 84,256 | |
| 1999 | Human Genome | 4,864,570 | 138,139 |
| 2000 | PubMed Central | 9,102,634 | 190,485 |
| 2001 | 13,602,262 | 233,977 | |
| 2002 | 19,808,101 | 283,590 | |
| 2003 | 29,819,397 | 380,104 | |
| 2004 | PubChem | 38,941,263 | 485,029 |
| 2005 | NIH Public Access | 49,152,445 | 666,917 |
| 2006 | Genome-Wide Association Studies | 62,765,195 | 864,586 |
| 2007 | Genome Reference Consortium | 77,632,813 | 958,584 |
| 2008 | 96,400,790 | 1,152,700 | |
| 2009 | 110,946,879 | 1,328,143 | |
| 2010 | 1000 Genomes | 125,764,384 | 1,501,335 |
| 2011 | 144,458,648 | 2,300,000 | |
| 2012 | Genetic Testing Registry and ClinVar | 157,889,737 | 2,430,000 |
| 2013 | MedGen and PubReader | 168,335,396 | 3,300,000 |
| 2014 | Pubmed Commons | 178,322,253 | 3,900,000 |
| 2015 | Food Pathogens Project | 188,372,017 | 4,000,000 |
| 2016 | 197,390,691 | 4,100,000 | |
| 2017 | 206,293,625 | 4,600,000 | |
| 2018 | Antimicrobial Resistance | 211,281,415 | 5,500,000 |
| 2019 | 215,333,020 | 6,200,000 |
NLM is the engine that powers health care data for clinical and observational research by driving intergovernmental activities and shaping the national agenda related to open science, AI, health data standards, and data science. NLM is working across the NIH and Federal Government, and in collaboration with the public and private sectors, to promote standards for exchanging biomedical research and clinical data to advance health care, public health, biomedical research, and product development. For example, in FY 2019, NLM played a critical role in the development, usage, and utility of a data exchange standard to improve flow and availability of data, the Health Level Seven International (HL7) Fast Healthcare Interoperability Resources (FHIR®)1. NIH is encouraging funded investigators to use the FHIR standard to capture, integrate, and exchange clinical data for research purposes and to enhance capabilities to share research data. NIH has also announced to the small business communities its special interest in supporting applications that use FHIR in the development of health IT products and services. To support these efforts, NLM is managing the development and testing of FHIR tools that researchers can use to increase the availability of high-quality, standardized research datasets and phenotypic information for genomic research and genomic medicine.
NLM also supports the development, maintenance and dissemination of health data standards and associated tools:
- NLM fosters standard coding systems for health care data elements that are used widely in health care and research, including LOINC® for clinical tests and measurements, SNOMED CT® for health conditions and other features, and RxNorm for clinical medications.
- NLM stewards the NIH Common Data Elements (CDE) Repository, a free, collaborative platform for sharing and discovering standard, structured, machine-readable definitions of data elements, standard variables, and measures used in NIH-funded clinical research. CDEs in the repository can be linked to existing health data standards and terminologies and can improve data quality and enable exchange and comparison of data across multiple studies and EHRs for clinical research and patient registries. In FY 2019, NLM updated and added new CDEs to the repository from six NIH ICs, including Patient-Reported Outcomes, PhenX Toolkit, preclinical traumatic brain injury, and Newborn Screening Translational Research Network. NLM also developed training materials, improved exports in standard formats, and supported data exchange with other NLM terminology systems. NLM also completed participation in two HHS interagency projects that drew on the informatics expertise of the NLM team and on the functionality of the NIH CDE Repository. In the last month of FY 2019, NLM launched a study to analyze user needs for CDEs and the NIH CDE Repository.
- NLM increases efficiency and accuracy of creating and maintaining value sets in the Value Set Authority Center (VSAC) by adding rule-based functionality (e.g., include all codes that are descendants of the SNOMED CT code for “Opioid dependence (disorder)”. VSAC enables EHR developers and biomedical researchers to download lists of codes and corresponding terms (i.e., value sets) that define clinical concepts used in clinical quality measures (e.g., patients with diabetes, tricyclic antidepressants).
- NLM provides comprehensive information on medication content and labeling for over 110,000 drugs marketed in the United States, with mobile access through DailyMed.
- NLM provides information on 240,000 implantable devices through NLM’s AccessGUDID portal, which facilitates machine-readable access to the FDA’s Global Unique Device Identification Database (GUDID).
In addition, NLM invests in outreach and training to help researchers, clinicians, educators, NLM staff, and the public use NLM’s wide range of biomedical information services. NLM’s exhibitions, traveling banner displays and web-based content reach underserved populations and promote interest in science, medicine, and technology. From FY 2016-2019, NLM’s traveling exhibitions reached more than 3 million visitors in 373 venues in 299 cities across the country, plus international locations reaching U.S. Armed Forces, through traveling exhibitions.
1 https://nlmdirector.nlm.nih.gov/2019/07/30/accelerating-innovation-in-science/
Budget Policy
The FY 2021 President’s Budget estimate for NLM’s Intramural Programs is $335.4 million, a decrease of $34.0 million from the FY2020 Enacted Budget of $369.4 million. NLM’s FY 2020 estimated expenditures for intramural research are approximately two-thirds of the total for NLM Intramural Programs and will remain unchanged in FY 2021. NLM’s intramural research program advances knowledge in data science methods, including AI and machine learning, as well as the advanced integration and analysis of genomic data, clinical research data, and observational health data. Consistent with recommendations of a recent Blue-Ribbon Panel review, NLM is aligning its intramural research activities under a single Scientific Director and seeking efficiencies in core research services. Research priorities will be informed by NLM’s needs for biomedical information services, including research in health data standards; computational biology; computational approaches to data curation; and machine-learning approaches to text and image processing.
NLM will support key intramural data science activities, including automated indexing and management of the biomedical literature, processing and organizing genomic data resulting from NIH-wide investments in whole genome sequencing, and genomic evaluations of pathogens in support outbreak detection and response by CDC, FDA, and USDA. The Library will prioritize and seek new efficiencies in its service offerings to ensure needed levels of support for biomedical and health information resources that are most heavily used by the scientific research community and the public and that support NIH-wide data science goals and data sharing policies. NLM will support further enhancements to ClinicalTrials.gov to facilitate submission of and access to clinical trial data submitted in accordance with the Food and Drug Administration Amendments Act of 2007 and NIH policy. It will improve tools for updating and disseminating clinical terminology standards required for interoperability of electronic health data and to support advanced integration and analysis of genomic, clinical research, and observational health data. In FY 2021, NLM will prioritize its outreach programs to improve efforts to promote access and training in the effective use of NLM resources, including data repositories, and to increase engagement with broad sets of stakeholders in the public and private sectors to leverage their talents, capabilities, and information resources. Through its cooperative agreement with the National Network of Libraries of Medicine, NLM will continue to support efforts to advance community engagement in the All of Us initiative
Extramural Programs
NLM’s extramural programs encompass three major activities: 1) Biomedical Informatics Research; 2) Health Information for Health Professionals and the Public; and 3) Informatics Resources for Biomedicine and Health. NLM is expanding its extramural programs to support growing demand for innovation in data science. In FY 2019, NLM funded 177 awards, including 26 that were co-funded with other NIH ICs.
Biomedical Informatics Research: NLM research awards in biomedical informatics aim to bring methods and concepts of biomedical informatics and data science to bear on problems in basic biomedical and behavioral research, health care, public health, consumer health, and other domains. NLM increased its budget for extramural research by $8.8 million between FY 2017 and FY 2019, representing a 60 percent growth in funding and almost doubling the number of Research Project Grant awards. NLM increased funding for early stage investigators from $5.3 million in FY 2018 to $6.2 million in FY 2019. NLM also funded 19 early-stage investigators in FY 2019, an increase from 10 in FY 2018.
NLM-funded researchers tested methods to capture, analyze, integrate, and curate biomedical data. NLM awarded research grants for projects including analytics for microbiome data, high-fidelity curation of gene-drug-phenotype relationships, and a data science-enabled approach to better manage endometriosis. For example, in FY 2019, NLM-funded researchers:
- Applied mathematical modeling and machine learning to patient data and physiology knowledge for personalized forecasts to help intensive care clinicians manage glucose.
- Applied computer vision and AI methods to images of neighborhood streets to create a “neighborhood looking glass” that pinpoints features of the built environment such as streets, parks, food stores, and health care facilities, which can affect health and health outcomes. This approach is being used to predict obesity and drug abuse in certain environments.
- Improved the effectiveness of clinically useful risk prediction approaches by applying a random field framework to match high-dimensional genomic data to many types of clinical data for predicting risks. This approach aims to use human genome data in the sphere of clinical decision-making. These techniques were used to investigate heterogeneous effects of 26 nicotine addiction-related genes and found strong associations, including a gender effect.
- Integrated clinical and genomic data about a patient matched to descriptions of open clinical trials and employed natural language processing to make it easy for clinical trialists to define characteristics of patients, thus helping them to optimize trial design.
- Applied machine learning and developed ‘digital phenotypes’ from patient data in EHRs to support ‘patient matching’ that allows a clinician to explore effective treatments for other similar patients, supporting decisions at the point of clinical care.
- Used unsupervised clustering, a machine learning technique, to mine massive amounts of high-throughput RNA-sequence data. Developed a computer program to quantify the degree to which cell types replicate across datasets. Results can help researchers find clusters of novel cell subtypes to identify candidate genes linked to organ development and disease.
NLM sparked innovative work in mission critical areas, resulting in research that addresses health challenges and disparities, such as on the health implications of the impact of food advertising targeted to racial and ethnic populations, approaches for semi-automated curation of research data, and the development of personal health records to support the unique needs of youth emancipating from foster care. NLM updated its general statement of interest for research and small business grants to emphasize incorporation of AI approaches into tools for clinicians, researchers, and consumers; models of complex data, information visualization that enhances understanding and usability; and translational research that integrates and links data relating to a person’s health from many sources (e.g., clinical, personal health, environmental, and neighborhood). NLM issued a Notice of Special Interest seeking applications for novel approaches to reduce inherent bias or missing data in collections of personal health data, such as those drawn from EHRs in a hospital. NLM collaborates with other NIH ICs to stimulate informatics and data science research in areas where data science can address a specific biomedical or clinical research objective (e.g., biomarker identification for pain and opioid use).
Health Information for Health Professionals and the Public: NLM supports outreach and engagement via the NNLM. The NNLM engages more than 7,000 academic health science libraries, hospital and public libraries, and community organizations across the United States and its territories to improve access to health information for everyone, from citizens to clinicians, research investigators, and data scientists, and it ensures equal access for all health professionals. NLM engages with public libraries to promote health and digital literacy and increase competency with consumer health information within the public library workforce. In FY 2019, NNLM staff supported 275 outreach and engagement projects, encompassing 2,452 activities across the United States, reaching more than 64,698 people. With public libraries, a recent focus for NNLM, more than 130 projects were funded, resulting in more than 1,400 activities reaching more than 13,180 people across 45 U.S. states. NNLM collaborates with NIH’s All of Us Research Program to provide high-quality health information and educate consumers about precision medicine in communities often underrepresented in biomedical research. NNLM also trains health sciences librarians in research data management and data science (see Program Portrait 2, page 15), encourages citizen science through promotion and education, and improves access to high-quality biomedical information online through #CiteNLM Wikipedia Edit-a-thons.
Informatics Resources for Biomedicine and Health: NLM’s Information Resources to Reduce Health Disparities program awards support projects that bring useful, usable health information to health disparity populations and their health care providers. NLM issues new awards for this program every other year. NLM made six new awards in FY 2019, up from four in 2017, focusing on topics including health disparities of migrant and seasonal farmers, heart health information for communities at risk of parasitic Chagas disease, information to help communities stop the spread of antibiotic resistant diseases, and environmental health literacy in Appalachia. In addition, NLM’s Grants for Scholarly Works in Biomedicine and Health supports scholarly publications. NLM made three such awards in FY 2019, focused on biomedical terminologies, dichlorodiphenyltrichloroethane (DDT) myths and science, and the electronic patient. These resource grants encourage collaborations between Historically Black Colleges and Universities, other Minority Serving Institutions, and other biomedical informatics training sites.
NLM also supports 16 university-based training programs in biomedical informatics. Many of NLM’s predoctoral and postdoctoral training programs expand outreach to other NIH training programs, involve partnerships with minority-serving organizations and information schools, and provide summer research experiences for high-school and undergraduate students from underrepresented groups. More than half of NLM’s funded training programs developed curriculum materials and student exercises related to biomedical data science, and shared resources freely via GitHub (see Program Portrait 2, page 18).
Budget Policy
The FY 2021 President’s Budget estimate includes $60.7 million for NLM’s Extramural Programs, a decrease of $6.2 million from the FY 2020 Enacted level of $66.9 million. Data science and informatics research is fundamental to the sophisticated information systems used to store, manage, display, and analyze research and health data. NLM will continue to accept investigator-initiated applications through NIH parent-grant announcements, as well as applications submitted to NLM’s own funding announcements, which focus on novel, broadly applicable methods in biomedical informatics and data science, personal health libraries and digital curation. In FY 2021, NLM will apply reductions averaging 15 percent to all noncompeting grants, including resource grants, in order to reduce the impact of the reduced FY 2021 funding level on new awards. NLM expects to award 35 new research project grants and will aim to support early stage and new investigators at success rates comparable to those of established investigators submitting new applications. NLM will continue to support its unique resource grant programs, career transition programs, at reduced levels. It will continue to support its highly regarded university-based training programs, maintaining the number of these programs.
Research Management and Support
NLM’s Research Management and Support activities provide administrative, budgetary, communications, and logistical support for NLM programs to ensure strategic planning, and evaluation; regulatory compliance; policy development; international coordination; and partnerships with other Federal agencies, Congress, the private sector, and the public. NLM is streamlining its organizational and administrative structure to enhance collaborative leadership, innovation, and customer service.
Budget Policy
The FY 2021 President’s Budget estimate is $19.6 million, a decrease of $1.0 million, from the FY 2020 Enacted level of $20.6 million. RMS will support NLM-wide planning and evaluation, including implementation of NLM’s strategic plan. It will also support enhancement of NLM’s information systems security, policy development and administration functions, and improved coordination of trans-NLM and trans-NIH efforts in data science.
Conclusion
Through its growing research programs, heavily-used information systems, and public engagement activities, NLM supports biomedical research and public health. NLM enables researchers, clinicians, and the public to use the vast wealth of biomedical data to improve the health of the Nation.
Budget Authority By Object Class 1
(Dollars in Thousands)
| FY 2020 Enacted | FY 2021 President's Budget | FY 2021 +/- FY 2020 | |
|---|---|---|---|
| Total compensable workyears: | |||
| Full-time equivalent | 741 | 741 | 0 |
| Full-time equivalent of overtime and holiday hours | 1 | 1 | 0 |
| Average ES salary | $197 | $198 | $0 |
| Average GM/GS grade | 11.8 | 11.8 | 0.0 |
| Average GM/GS salary | $111 | $112 | $1 |
| Average salary, grade established by act of July 1, 1944 (42 U.S.C. 207) | $118 | $118 | $0 |
| Average salary of ungraded positions | $157 | $157 | $0 |
| OBJECT CLASSES | FY 2020 Enacted | FY 2021 President's Budget | FY 2021 +/- FY 2020 | |
|---|---|---|---|---|
| Personnel Compensation | ||||
| 11.1 | Full-Time Permanent | 44,612 | 45,125 | 513 |
| 11.3 | Other Than Full-Time Permanent | 45,672 | 46,198 | 525 |
| 11.5 | Other Personnel Compensation | 1,738 | 1,758 | 20 |
| 11.7 | Military Personnel | 127 | 131 | 3 |
| 11.8 | Special Personnel Services Payments | 1,884 | 1,905 | 22 |
| 11.9 | Subtotal Personnel Compensation | $94,034 | $95,117 | $1,083 |
| 12.1 | Civilian Personnel Benefits | 30,829 | 32,032 | 1,202 |
| 12.2 | Military Personnel Benefits | 59 | 61 | 2 |
| 13.0 | Benefits to Former Personnel | 0 | 0 | 0 |
| Subtotal Pay Costs | $124,922 | $127,210 | $2,287 | |
| 21.0 | Travel & Transportation of Persons | 1,155 | 1,082 | -73 |
| 22.0 | Transportation of Things | 94 | 88 | -6 |
| 23.1 | Rental Payments to GSA | 0 | 0 | 0 |
| 23.2 | Rental Payments to Others | 184 | 171 | -13 |
| 23.3 | Communications, Utilities & Misc. Charges | 440 | 410 | -30 |
| 24.0 | Printing & Reproduction | 141 | 131 | -10 |
| 25.1 | Consulting Services | 67,464 | 55,880 | -11,584 |
| 25.2 | Other Services | 63,383 | 51,290 | -12,093 |
| 25.3 | Purchase of goods and services from government accounts | 66,447 | 65,402 | -1,045 |
| 25.4 | Operation & Maintenance of Facilities | 19,202 | 15,365 | -3,837 |
| 25.5 | R&D Contracts | 17 | 18 | 0 |
| 25.6 | Medical Care | 0 | 0 | 0 |
| 25.7 | Operation & Maintenance of Equipment | 16,646 | 13,527 | -3,119 |
| 25.8 | Subsistence & Support of Persons | 13 | 13 | 0 |
| 25.0 | Subtotal Other Contractual Services | $233,172 | $201,494 | -$31,678 |
| 26.0 | Supplies & Materials | 1,249 | 1,095 | -154 |
| 31.0 | Equipment | 28,654 | 23,277 | -5,378 |
| 32.0 | Land and Structures | 0 | 0 | 0 |
| 33.0 | Investments & Loans | 0 | 0 | 0 |
| 41.0 | Grants, Subsidies & Contributions | 66,897 | 60,705 | -6,191 |
| 42.0 | Insurance Claims & Indemnities | 0 | 0 | 0 |
| 43.0 | Interest & Dividends | 2 | 2 | 0 |
| 44.0 | Refunds | 0 | 0 | 0 |
| Subtotal Non-Pay Costs | $331,989 | $288,455 | -$43,533 | |
| Total Budget Authority by Object Class | $456,911 | $415,665 | -$41,246 | |
1 Includes FTEs whose payroll obligations are supported by the NIH Common Fund.
Salaries and Expenses
Dollars in Thousands
| OBJECT CLASSES | FY 2020 Enacted | FY 2021 President's Budget | FY 2021 +/- FY 2020 |
|---|---|---|---|
| Personnel Compensation | |||
| Full-Time Permanent (11.1) | $44,612 | $45,125 | $513 |
| Other Than Full-Time Permanent (11.3) | 45,672 | 46,198 | 525 |
| Other Personnel Compensation (11.5) | 1,738 | 1,758 | 20 |
| Military Personnel (11.7) | 127 | 131 | 3 |
| Special Personnel Services Payments (11.8) | 1,884 | 1,905 | 22 |
| Subtotal Personnel Compensation (11.9) | $94,034 | $95,117 | $1,083 |
| Civilian Personnel Benefits (12.1) | $30,829 | $32,032 | $1,202 |
| Military Personnel Benefits (12.2) | 59 | 61 | 2 |
| Benefits to Former Personnel (13.0) | 0 | 0 | 0 |
| Subtotal Pay Costs | $124,922 | $127,210 | $2,287 |
| Travel & Transportation of Persons (21.0) | $1,155 | $1,082 | -$73 |
| Transportation of Things (22.0) | 94 | 88 | -6 |
| Rental Payments to Others (23.2) | 184 | 171 | -13 |
| Communications, Utilities & Misc. Charges (23.3) | 440 | 410 | -30 |
| Printing & Reproduction (24.0) | 141 | 131 | -10 |
| Other Contractual Services: | |||
| Consultant Services (25.1) | 67,464 | 55,880 | -11,584 |
| Other Services (25.2) | 63,383 | 51,290 | -12,093 |
| Purchases from government accounts (25.3) | 55,558 | 52,771 | -2,787 |
| Operation & Maintenance of Facilities (25.4) | 19,202 | 15,365 | -3,837 |
| Operation & Maintenance of Equipment (25.7) | 16,646 | 13,527 | -3,119 |
| Subsistence & Support of Persons (25.8) | 13 | 13 | 0 |
| Subtotal Other Contractual Services | $222,266 | $188,846 | -$33,421 |
| Supplies & Materials (26.0) | $1,249 | $1,095 | -$154 |
| Subtotal Non-Pay Costs | $225,529 | $191,823 | -$33,707 |
| Total Administrative Costs | $350,452 | $319,032 | -$31,420 |
Detail of Full-Time Equivalent Employment (FTE)
| OFFICE/DIVISION | FY 2019 Final | FY 2020 Enacted | FY 2021 President's Budget | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Civilian | Military | Total | Civilian | Military | Total | Civilian | Military | Total | |
| Division of Extramural Programs | |||||||||
| Direct: | 18 | - | 18 | 19 | - | 19 | 19 | - | 19 |
| Reimbursable: | - | - | - | - | - | - | - | - | - |
| Total: | 18 | - | 18 | 19 | - | 19 | 19 | - | 19 |
| Division of Library Operations | |||||||||
| Direct: | 243 | - | 243 | 291 | - | 291 | 291 | - | 291 |
| Division of Library Operations | |||||||||
| Reimbursable: | - | - | - | - | - | - | - | - | - |
| Division of Library Operations | |||||||||
| Total: | 243 | - | 243 | 291 | - | 291 | 291 | - | 291 |
| Division of Specialized Information Services | |||||||||
| Direct: | 20 | - | 20 | - | - | - | - | - | - |
| Reimbursable: | - | - | - | - | - | - | - | - | - |
| Total: | 20 | - | 20 | - | - | - | - | - | - |
| Lister Hill National Center for Biomedical Communications | |||||||||
| Direct: | 40 | - | 40 | 45 | - | 45 | 45 | - | 45 |
| Reimbursable: | - | - | - | - | - | - | - | - | - |
| Total: | 40 | - | 40 | 45 | - | 45 | 45 | - | 45 |
| National Center for Biotechnology Information | |||||||||
| Direct: | 270 | 1 | 271 | 300 | 1 | 301 | 300 | 1 | 301 |
| Reimbursable: | 1 | - | 1 | 1 | - | 1 | 1 | - | 1 |
| Total: | 271 | 1 | 272 | 301 | 1 | 302 | 301 | 1 | 302 |
| Office of the Director/Administration | |||||||||
| Direct: | 53 | - | 53 | 71 | - | 71 | 71 | - | 71 |
| Reimbursable: | 13 | - | 13 | 13 | - | 13 | 13 | - | 13 |
| Total: | 66 | - | 66 | 84 | - | 84 | 84 | - | 84 |
| Total | 658 | 1 | 659 | 740 | 1 | 741 | 740 | 1 | 741 |
Includes FTEs whose payroll obligations are supported by the NIH Common Fund.
| OFFICE/DIVISION | FY 2019 Final | FY 2020 Enacted | FY 2021 President's Budget | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Civilian | Military | Total | Civilian | Military | Total | Civilian | Military | Total | |
| FTEs supported by funds from Cooperative Research and Development Agreements. | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| FISCAL YEAR | Average GS Grade |
|---|---|
| 2017 | 11.5 |
| 2018 | 11.9 |
| 2019 | 11.8 |
| 2020 | 11.8 |
| 2021 | 11.8 |
Detail of Positions 1
| GRADE | FY 2019 Final | FY 2020 Enacted | FY 2021 President's Budget |
|---|---|---|---|
| Total, ES Positions | 5 | 5 | 5 |
| Total, ES Salary | 958,665 | 987,425 | 987,820 |
| GM/GS-15 | 17 | 17 | 17 |
| GM/GS-14 | 46 | 46 | 46 |
| GM/GS-13 | 121 | 146 | 146 |
| GS-12 | 111 | 138 | 138 |
| GS-11 | 21 | 21 | 21 |
| GS-10 | 0 | 0 | 0 |
| GS-9 | 12 | 12 | 12 |
| GS-8 | 32 | 32 | 32 |
| GS-7 | 6 | 6 | 6 |
| GS-6 | 1 | 1 | 1 |
| GS-5 | 2 | 2 | 2 |
| GS-4 | 1 | 1 | 1 |
| GS-3 | 3 | 3 | 3 |
| GS-2 | 5 | 5 | 5 |
| GS-1 | 2 | 2 | 2 |
| Subtotal | 380 | 432 | 432 |
| Grades established by Act of July 1, 1944 (42 U.S.C. 207) | |||
| Assistant Surgeon General | 0 | 0 | 0 |
| Director Grade | 0 | 0 | 0 |
| Senior Grade | 1 | 1 | 1 |
| Full Grade | 0 | 0 | 0 |
| Senior Assistant Grade | 0 | 0 | 0 |
| Assistant Grade | 0 | 0 | 0 |
| Subtotal | 1 | 1 | 1 |
| Ungraded | 271 | 303 | 303 |
| Total permanent positions | 375 | 457 | 457 |
| Total positions, end of year | 657 | 741 | 741 |
| Total full-time equivalent (FTE) employment, end of year | 659 | 741 | 741 |
| Average ES salary | 191,733 | 197,485 | 197,564 |
| Average GM/GS grade | 11.8 | 11.8 | 11.8 |
| Average GM/GS salary | 107,637 | 110,866 | 111,975 |
1 Includes FTEs whose payroll obligations are supported by the NIH Common Fund.
Last Reviewed: February 10, 2026

